models — Fitting models#
This page documents the various fitting models readily available in the package.
Polynomials#
Constant#
A constant model. (Note this is equivalent to a polynomial of degree zero, but we keep it separate for clarity.)
Fitting with a constant model is equivalent to computing the weighted average of the values of the dependent variable.
Line#
A straight-line model. (Note this is equivalent to a polynomial of degree one, but we keep it separate for clarity.)
Polynomial#
A simple polynomial model of arbitrary degree.
The degree of the polynomial is set at initialization time via the degree argument
in the constructor, e.g.,
>>> quadratic = Polynomial(2)
>>> cubic = Polynomial(degree=3)
Quadratic#
Simple alias for a polynomial of degree two.
Cubic#
Simple alias for a polynomial of degree three.
Exponentials and power-laws#
PowerLaw#
A power-law model with the general form:
Note
The overloaded plot() method automatically switches to a log-log scale.
Exponential#
A simple exponential model:
Note
Note that the location parameter is not a fit parameter, but rather a fixed
offset that is set when the model instance is created, e.g.,
>>> exponential = Exponential(location=2.0)
The basic idea behind this is to avoid the degeneracy between the location and the prefactor, and this is one of the main reasons this model is not implemented wrapping the scipy exponential distribution.
ExponentialComplement#
The exponential complement, describing an exponential rise:
Note
See notes on Exponential regarding the location parameter.
StretchedExponential#
A stretched exponential model:
Note
See notes on Exponential regarding the location parameter.
StretchedExponentialComplement#
The complement of the stretched exponential model:
Note
See notes on Exponential regarding the location parameter.
Sigmoid models#
Sigmoid models are location-scale models defined in terms of a standardized shape function \(g(z)\)
where \(A\) is the amplitude (total height of the sigmoid), \(m\) is the location (typically the point where the value of the function if 50% of the amplitude) and \(s\) is the scale parameter, representing the width of the transtion.
Note
In this case the amplitude parameter does not represent an area (as in peak-like models), but rather the total increase of the function from its lower asymptote to its upper asymptote.
Note when the scale parameter is negative, we switch to the complement of the sigmoid function, i.e., a monotonically decreasing function from 1 to 0 (in standard form).
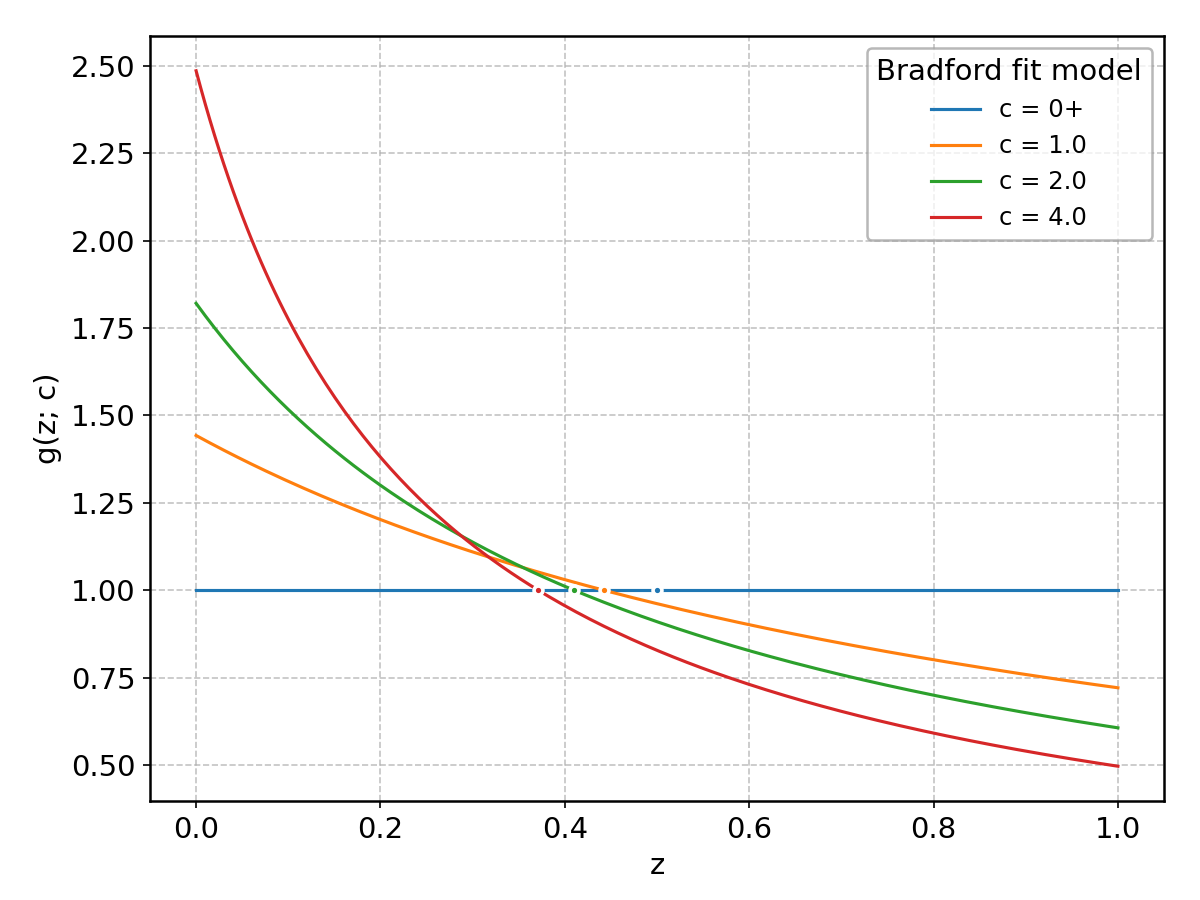
\(g(z)\) is generally a monotonically increasing function, ranging from 0 to 1 as its argument goes from -infinity to +infinity, as illustrated below for some of the models available in the package.
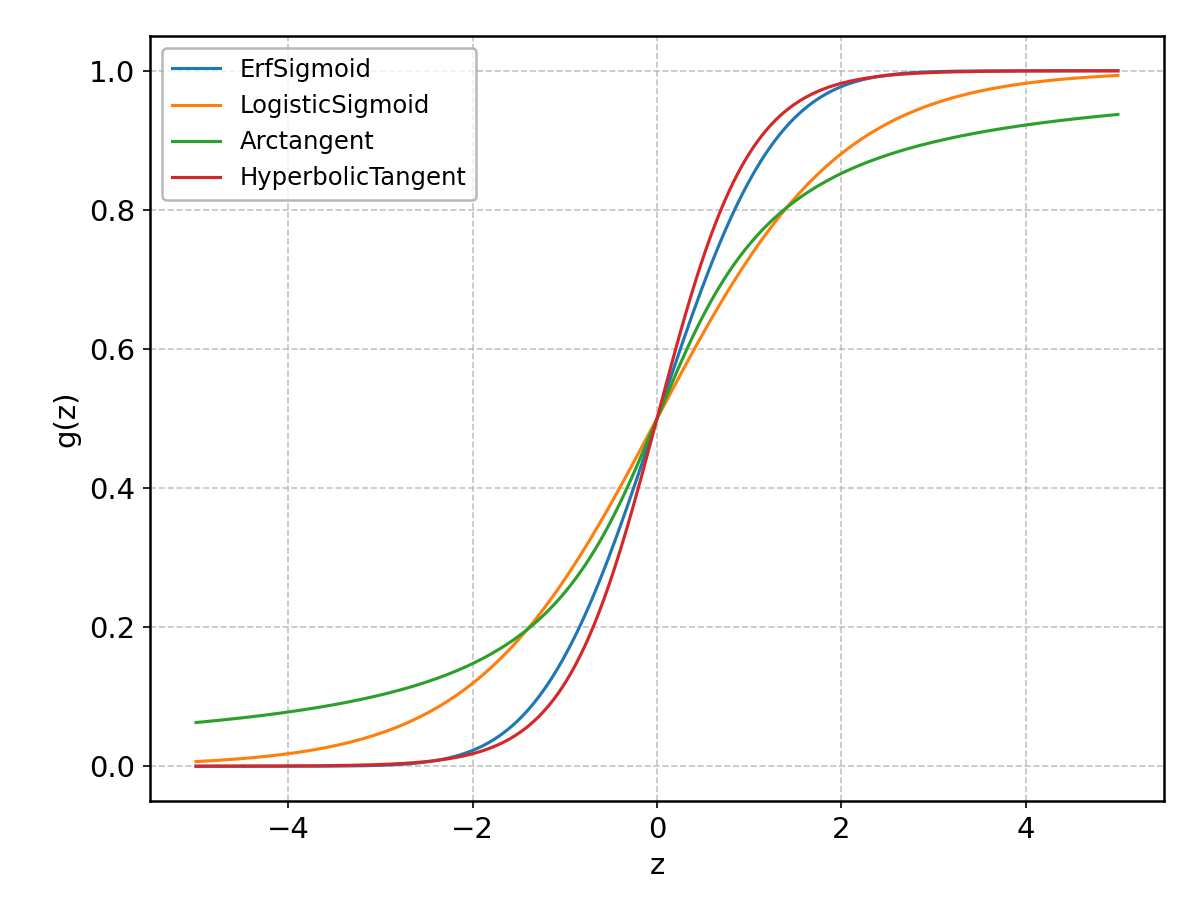
ErfSigmoid#
The cumulative function of a gaussian distribution:
Warning
The naming might be slightly unfortunate here, as, strictly speaking, this is not
the error function defined, e.g., in scipy.special, but hopefully it is clear
enough in the context of model fitting.
LogisticSigmoid#
A logistic sigmoid defined by the standard shape function:
Arctangent#
An arctangent sigmoid defined by the standard shape function:
@staticmethod
def shape(z):
# pylint: disable=arguments-differ
return 0.5 + np.arctan(z) / np.pi
HyperbolicTangent#
An hyperbolic tangent sigmoid defined by the standard shape function:
Continuous random variables#
Most of the models defined in this package are wrappers around continuous random
variables defined in scipy.stats, which provide a large variety of
standardized distributions. Most (but not all) of these distributions can be
interpreted as peak-like models. All of them are location-scale models
defined in terms of a standardized shape function \(g(z)\)
where \(A\) is the amplitude (area under the peak), \(m\) is the location
(parameter specifying the peak position), and \(s\) is the scale (parameter
specifying the peak width). The trailing \ldots indicates any additional shape
parameters that might be required by the specific distribution.
See also
The rest of this section lists all the available distributions, with a brief
description of their support and shape parameters. For more details on each
distribution, please refer to the corresponding documentation in scipy.stats.
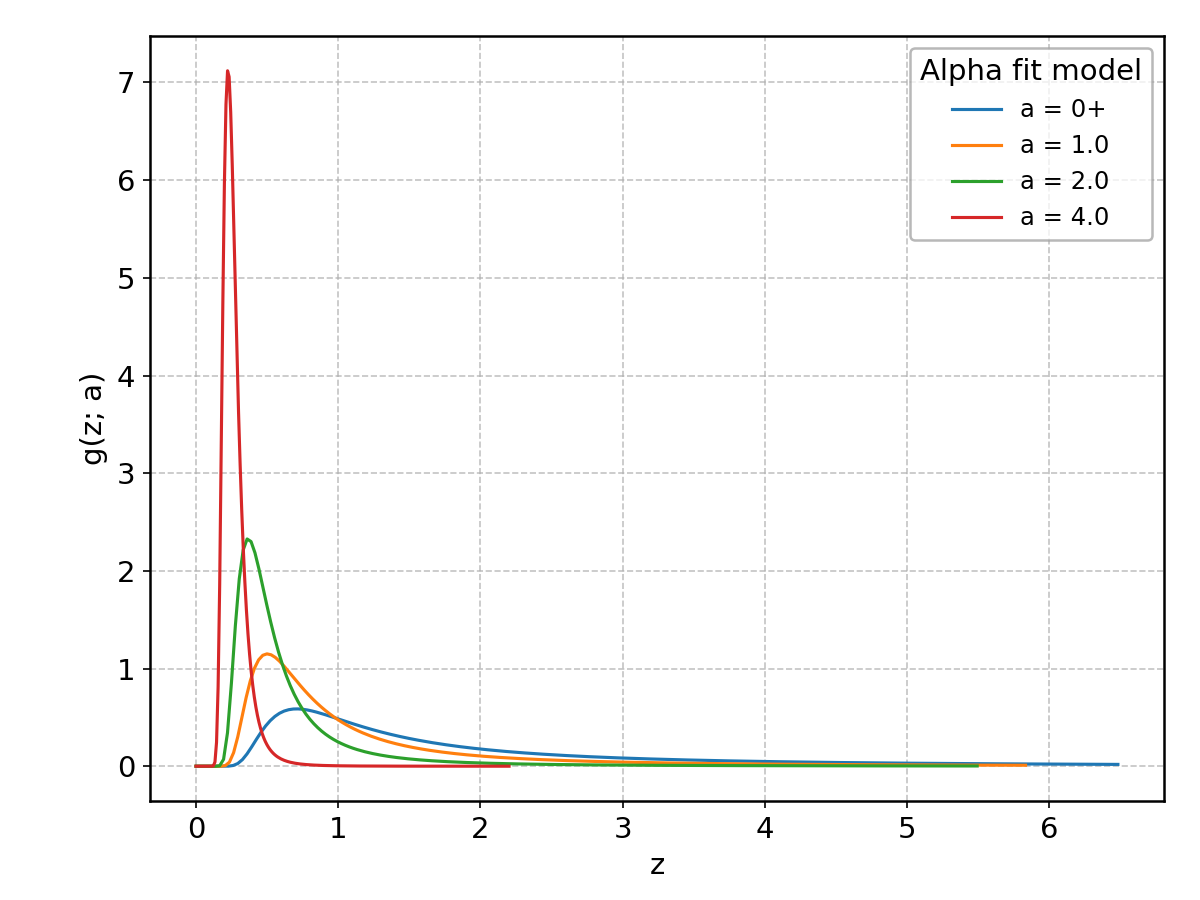
Alpha#
Wrapped from scipy.stats.alpha; support: \(z > 0\); shape parameter(s): \(a > 0\).
(Note the mean and the standard deviation of the distribution are always infinite.)
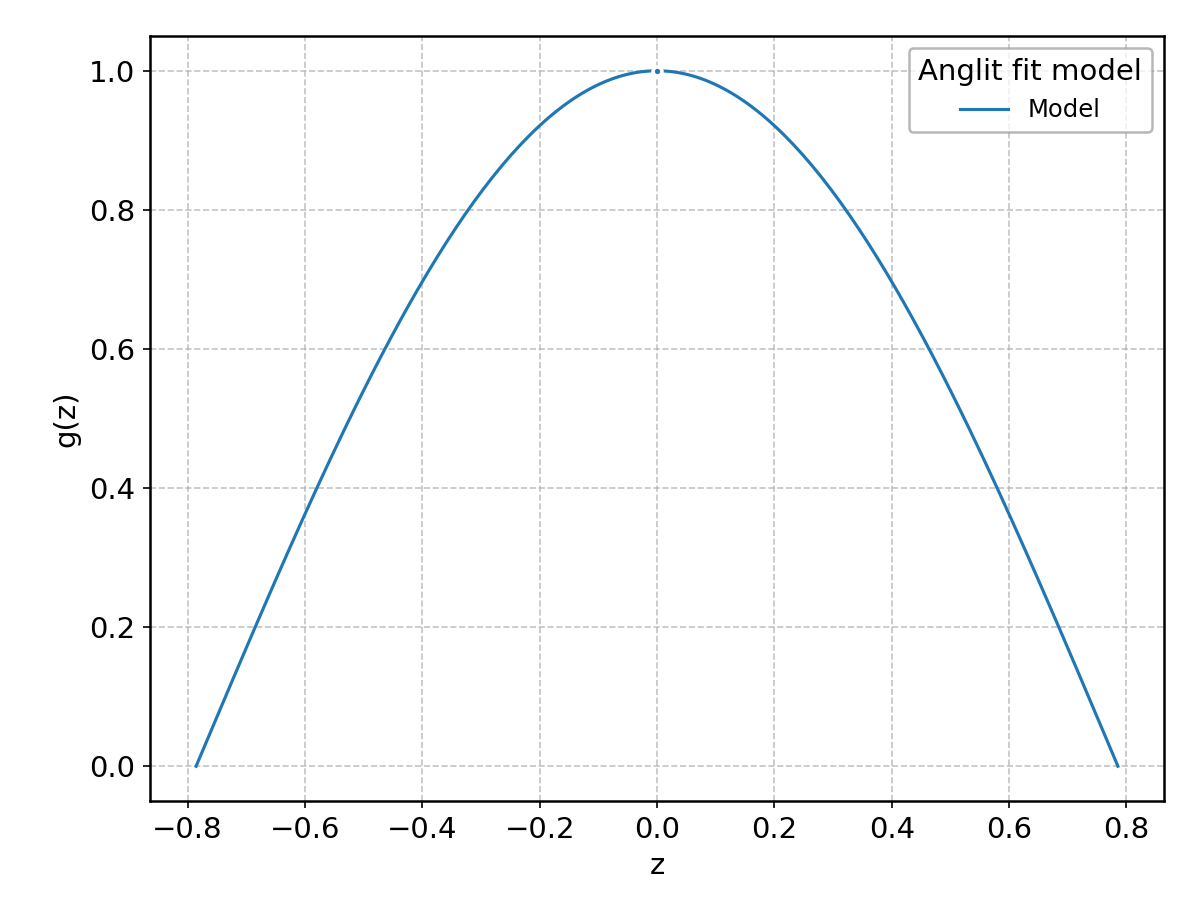
Anglit#
Wrapped from scipy.stats.anglit; support: \(-\pi/4 \le z \le \pi/4\).
Arcsine#
Wrapped from scipy.stats.arcsine; support: \(0\le z \le 1\).
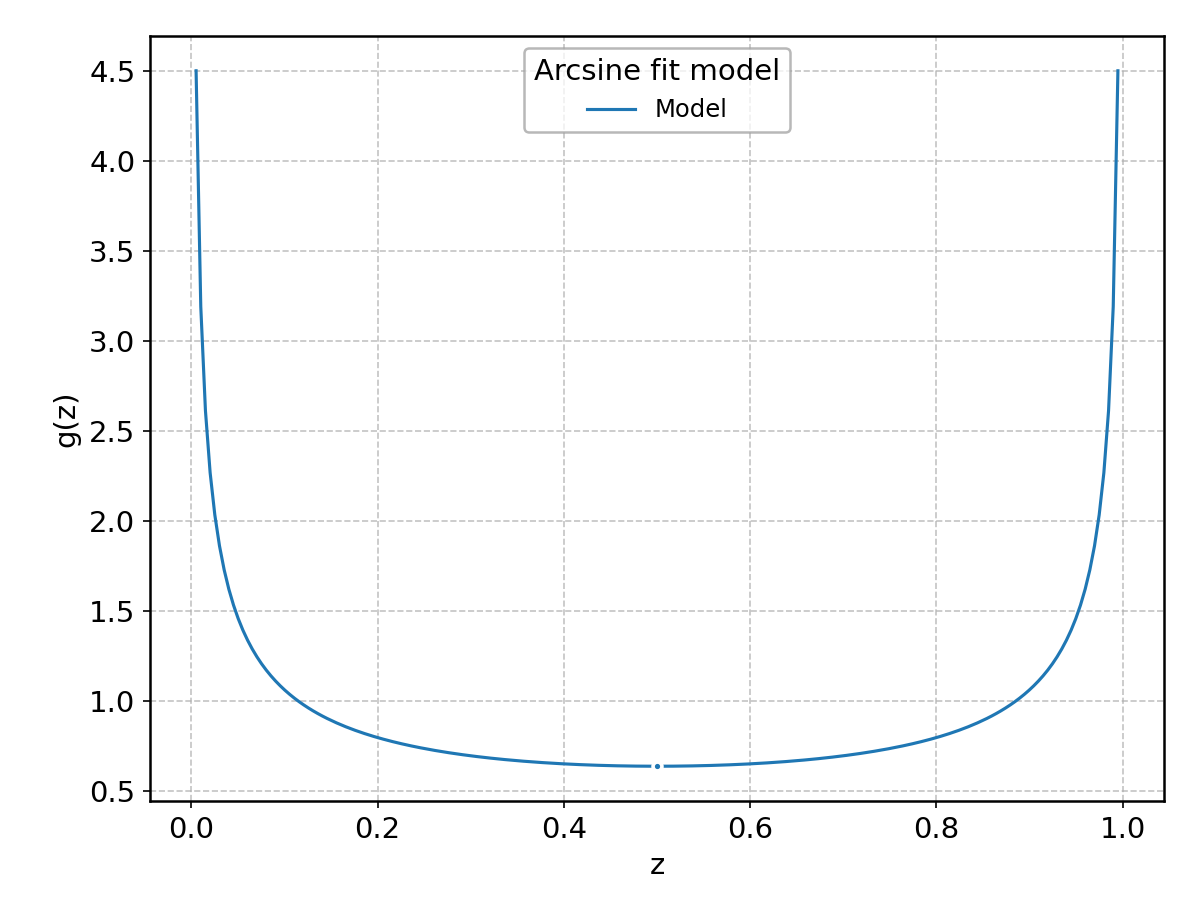
Argus#
Wrapped from scipy.stats.argus; support: \(0 < z < 1\); shape parameter(s): \(\chi > 0\).
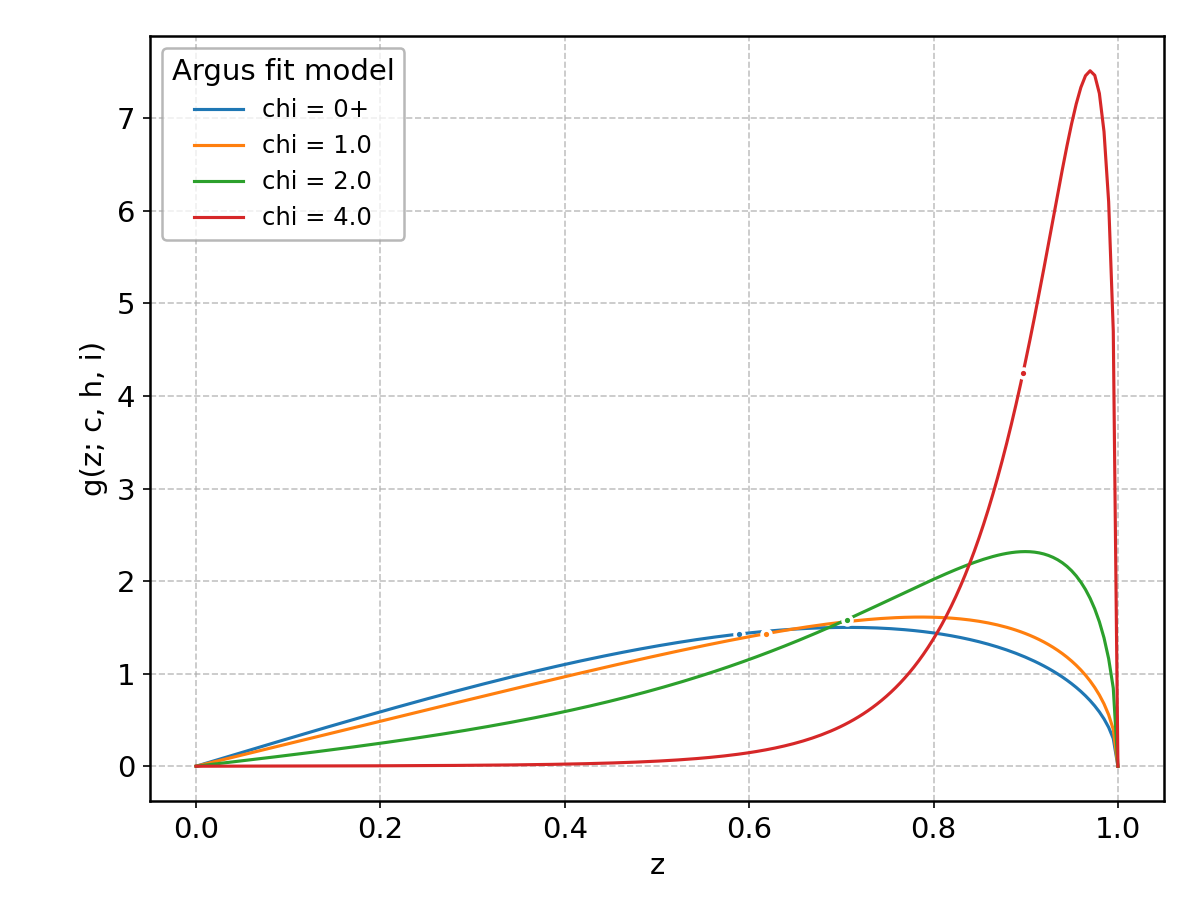
Beta#
Wrapped from scipy.stats.beta; support: \(0 < z < 1\); shape parameter(s): \(a> 0\), \(b > 0\).
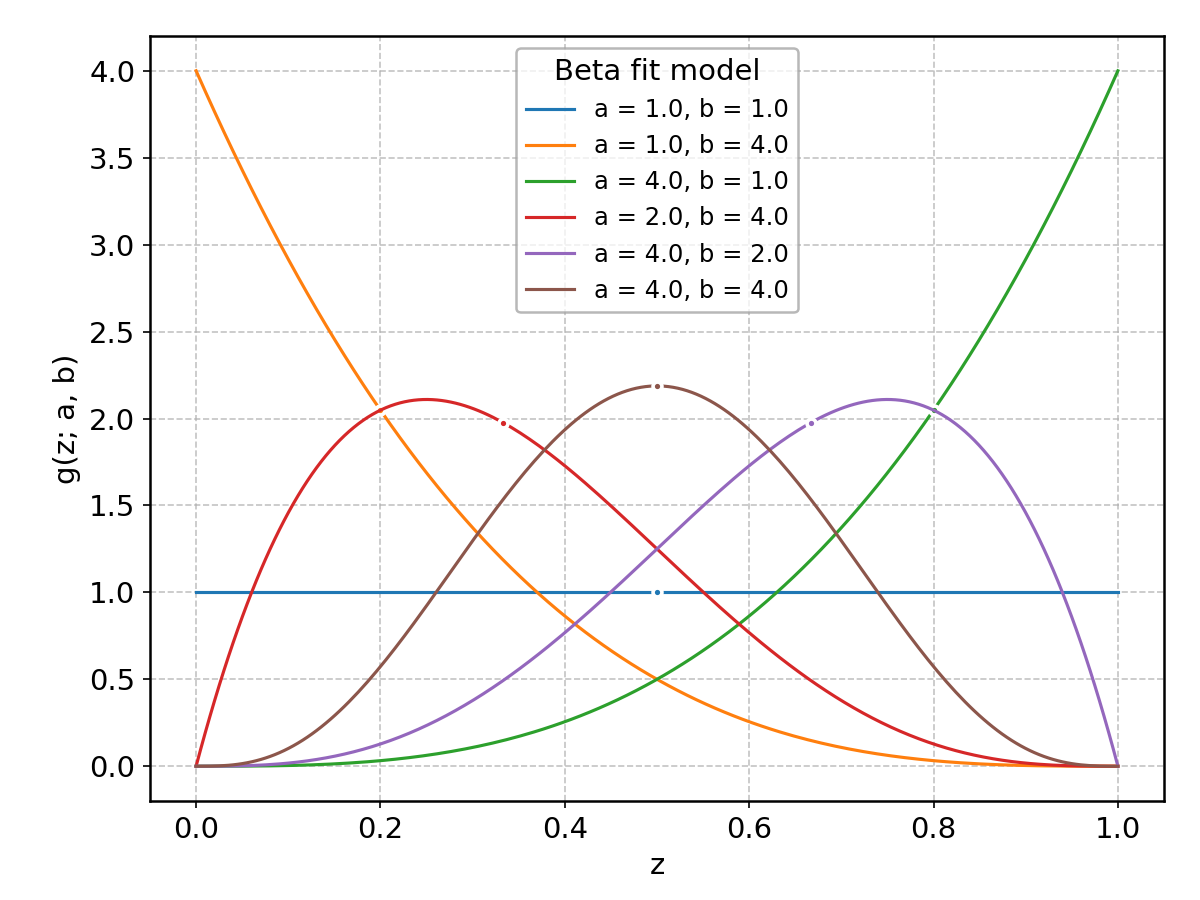
BetaPrime#
Wrapped from scipy.stats.betaprime; support: \(0 < z < 1\); shape parameter(s): \(a> 0\), \(b > 0\).
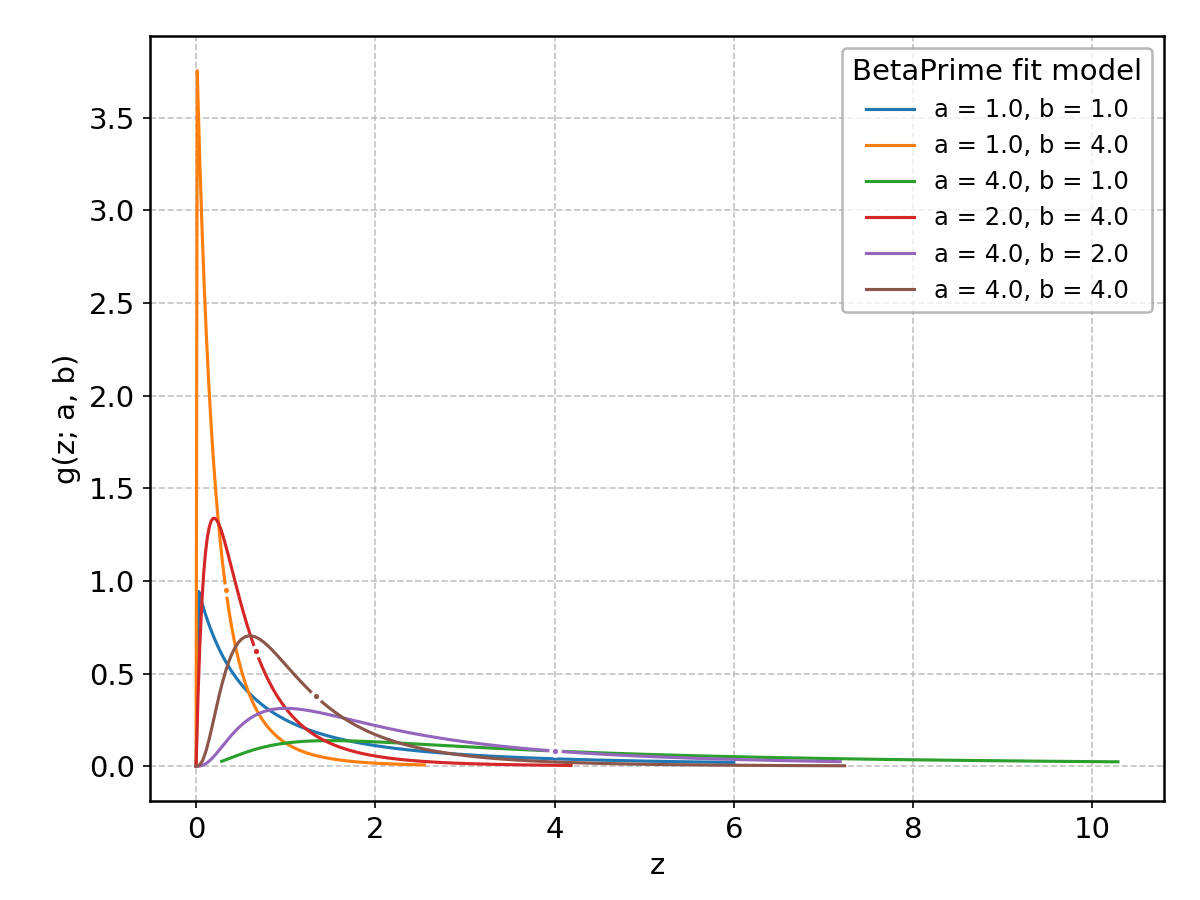
Bradford#
Wrapped from scipy.stats.bradford; support: \(0 < z < 1\); shape parameter(s): \(c > 0\).
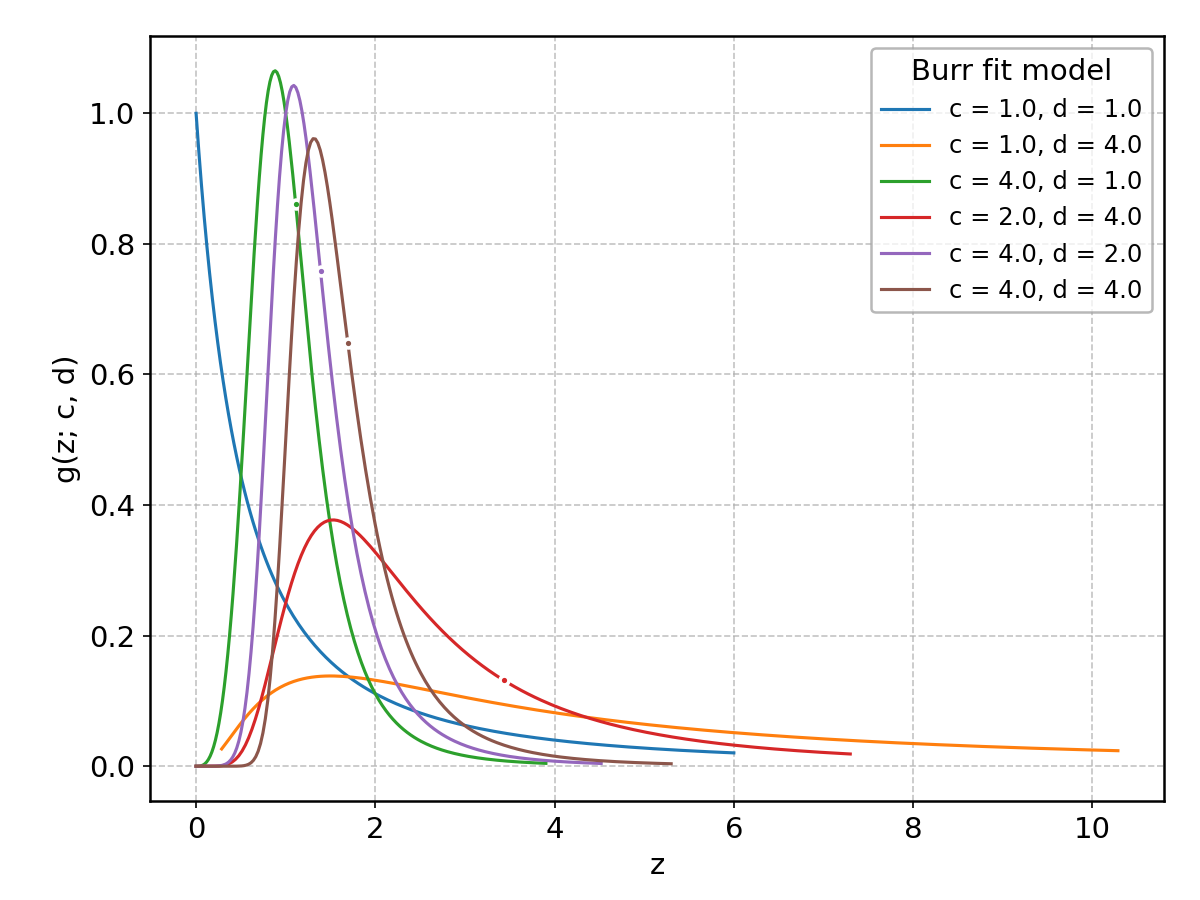
Burr#
Wrapped from scipy.stats.burr; support: \(z > 0\); shape parameter(s): \(c, d > 0\).
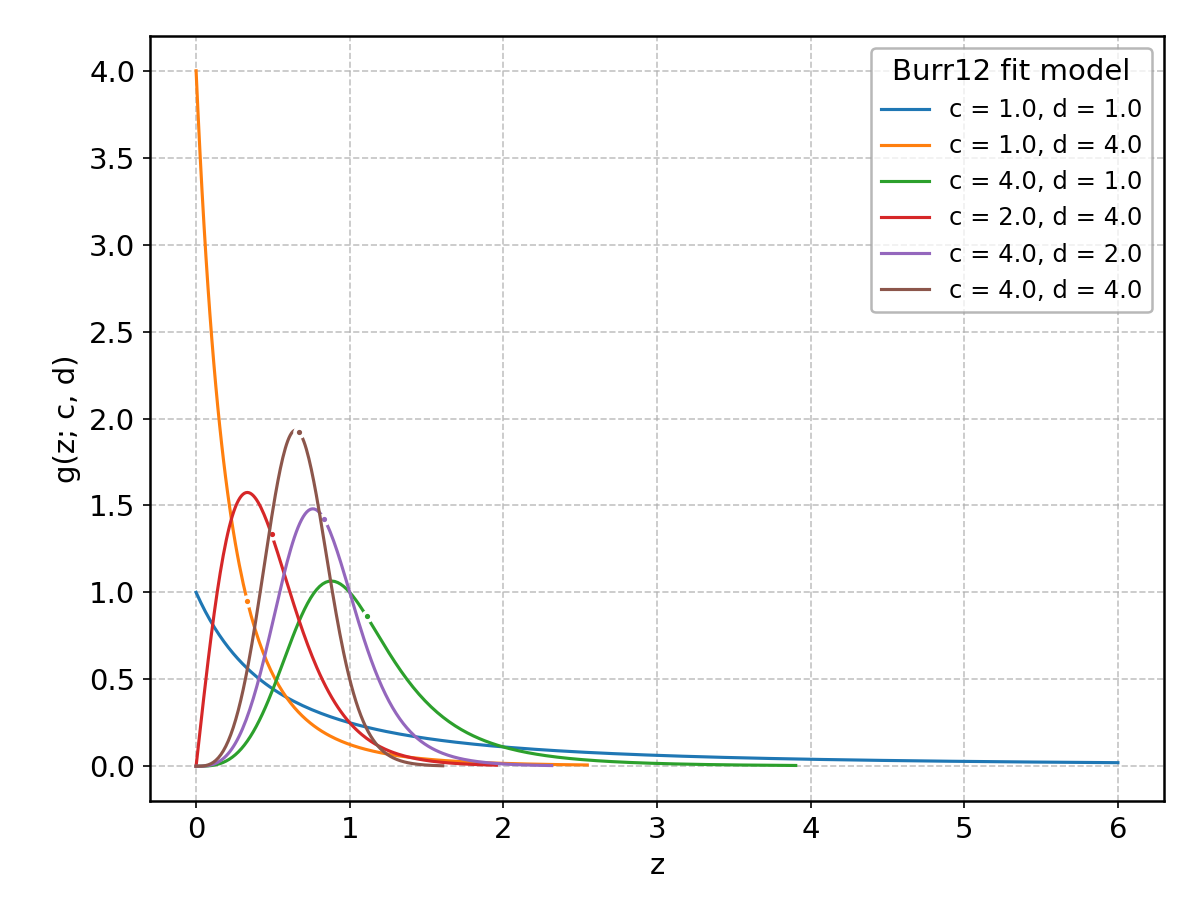
Burr12#
Wrapped from scipy.stats.burr12; support: \(z > 0\); shape parameter(s): \(c, d > 0\).
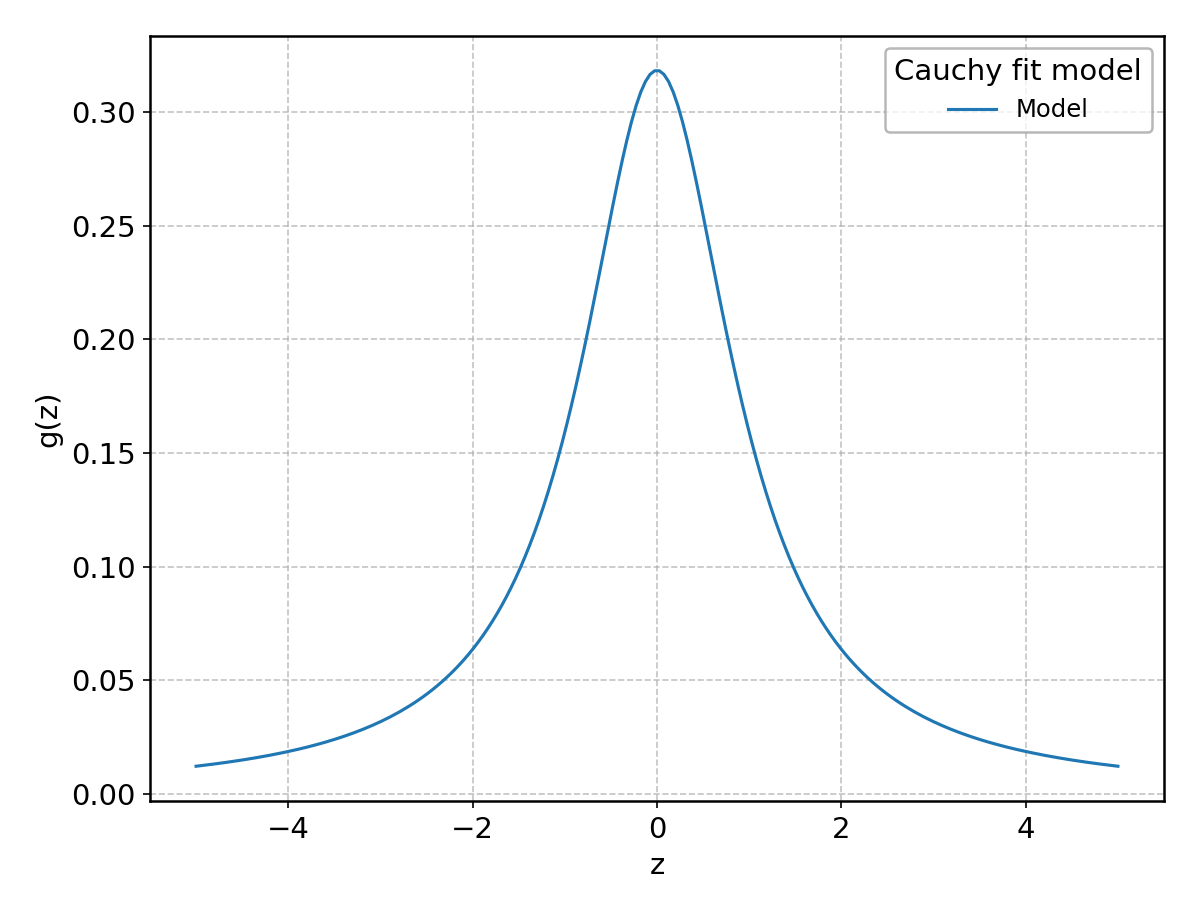
Cauchy#
Wrapped from scipy.stats.cauchy; support: \(z > 0\).
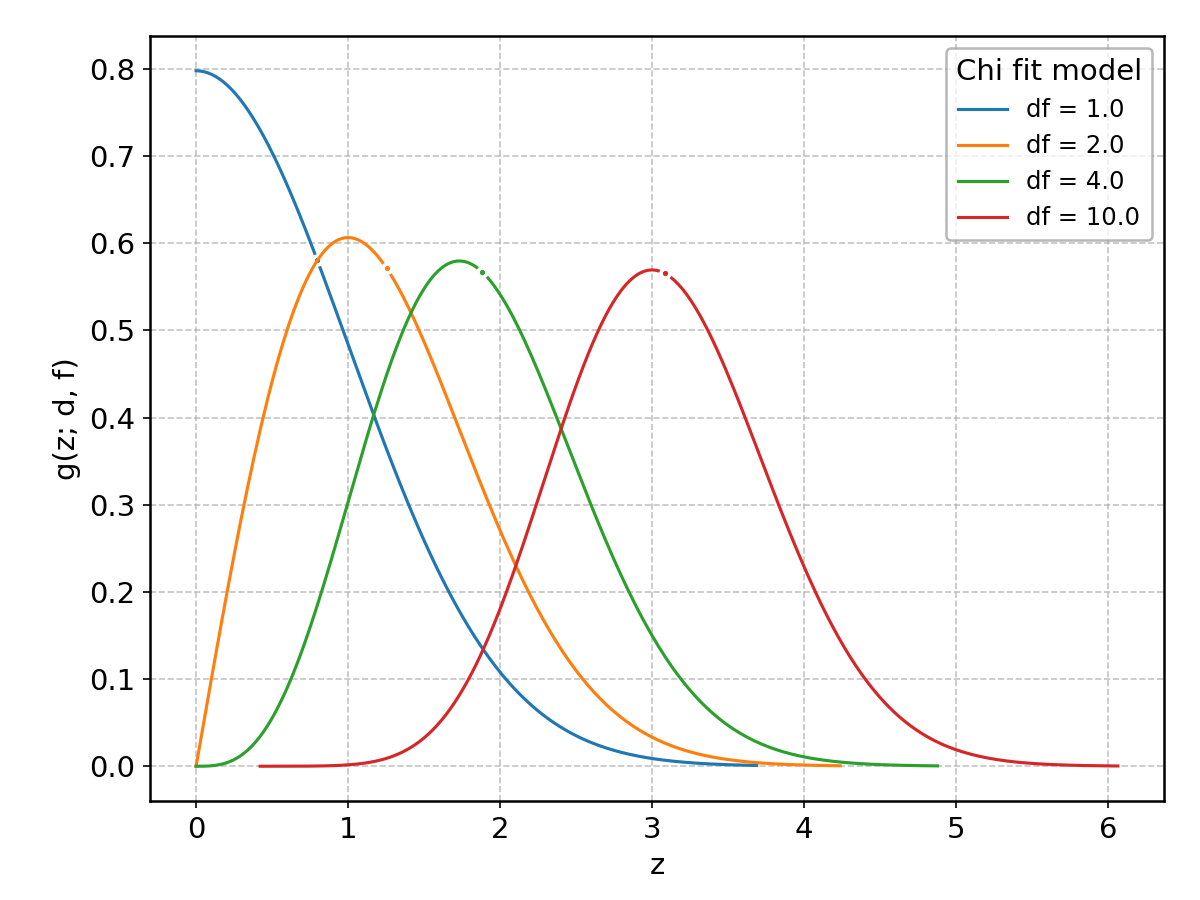
Chi#
Wrapped from scipy.stats.chi; support: \(z > 0\); shape parameter(s): \(\text{df} > 0\).
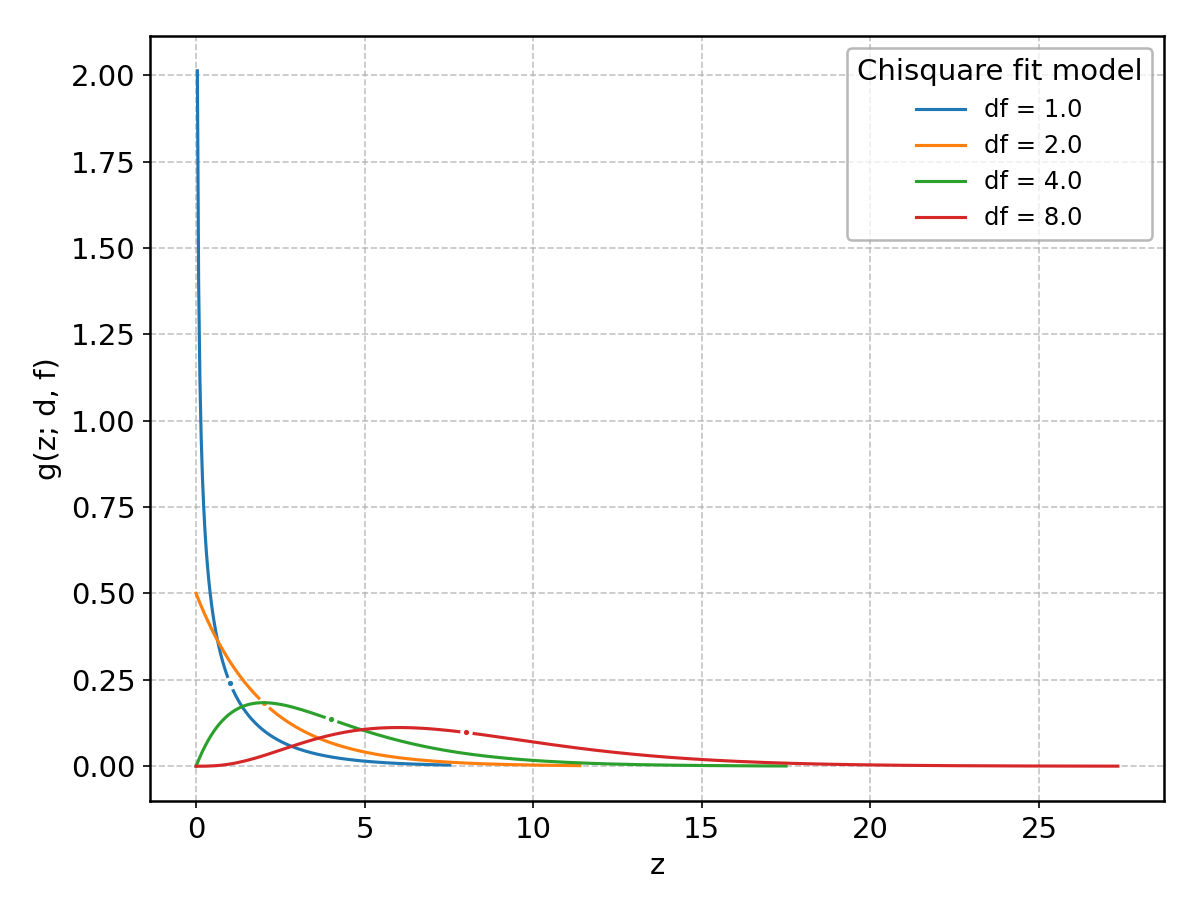
Chisquare#
Wrapped from scipy.stats.chi2; support: \(z > 0\) shape parameter(s): \(\text{df} > 0\).
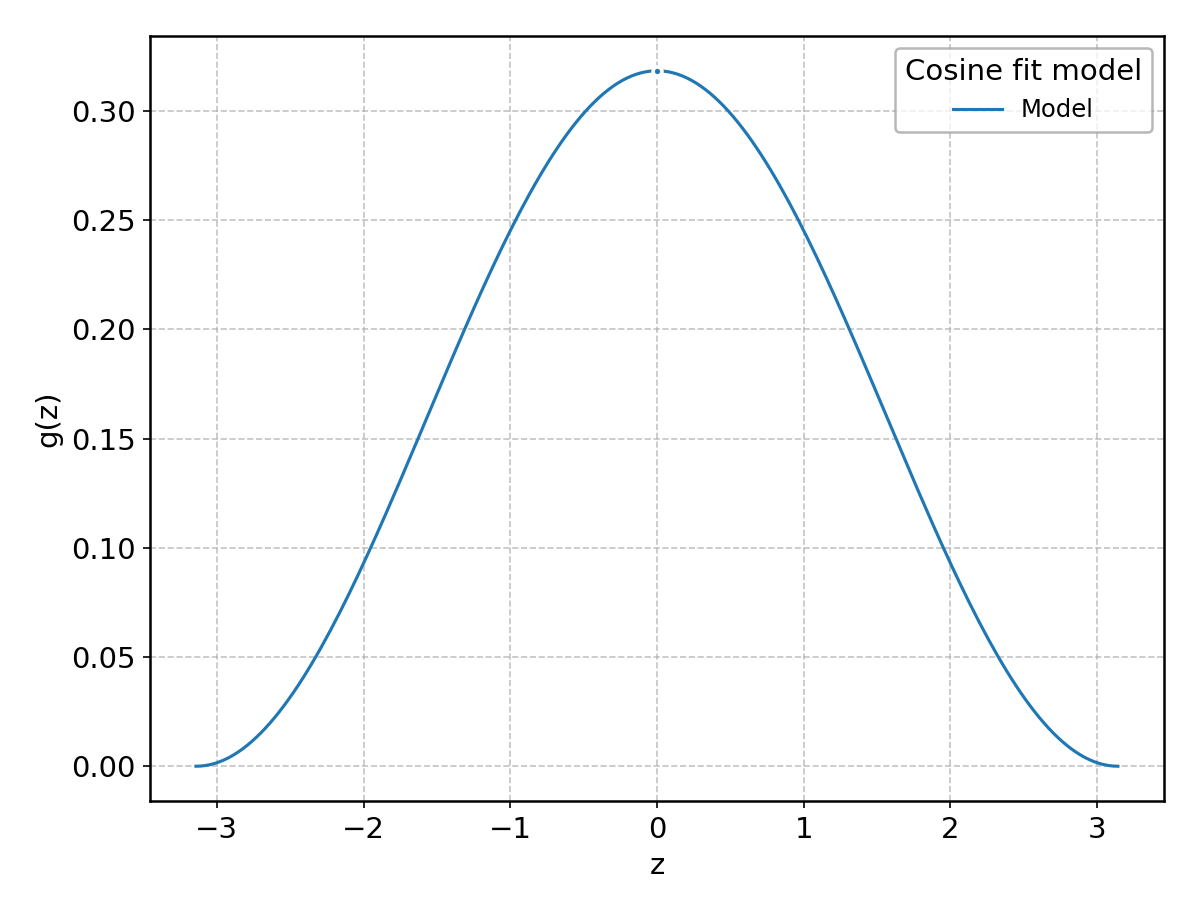
Cosine#
Wrapped from scipy.stats.cosine; support: \(-\pi \le z \le \pi\).
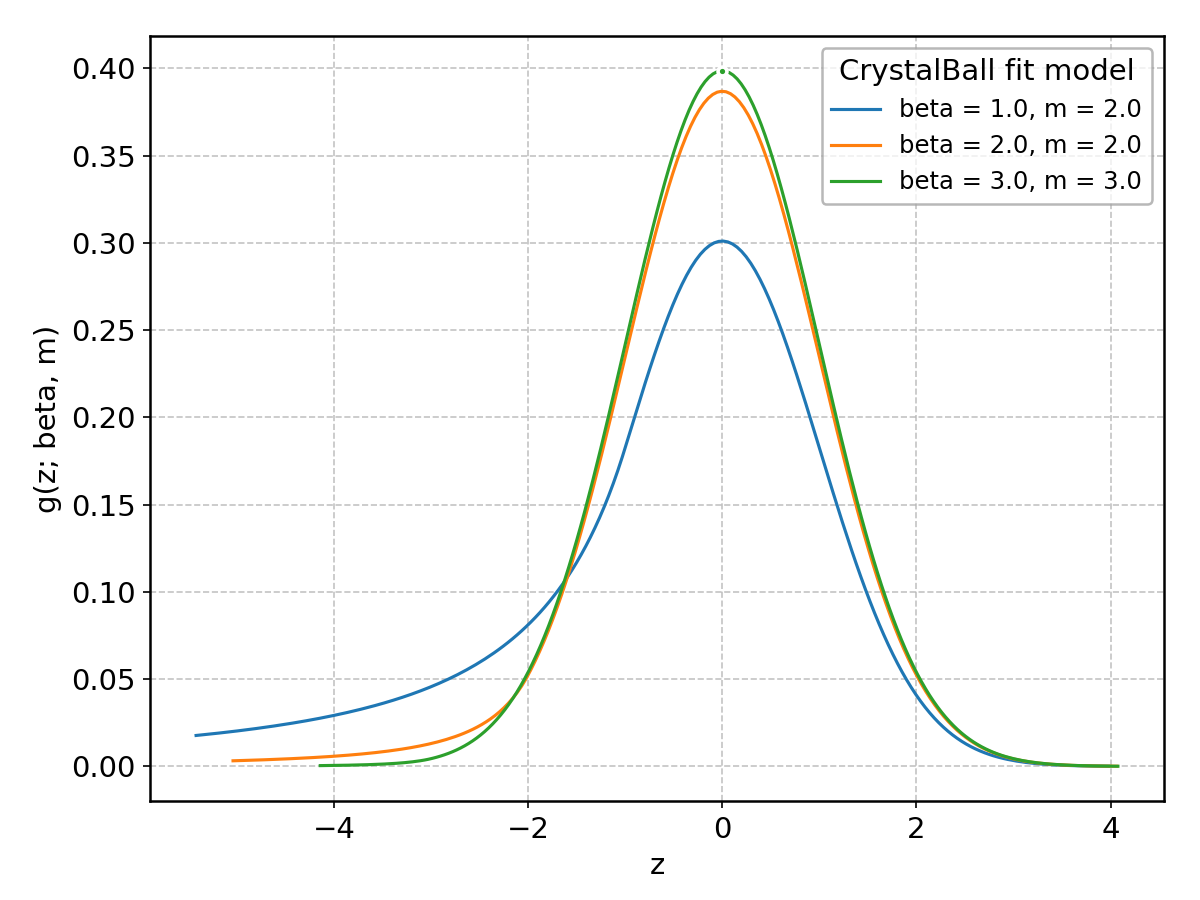
CrystalBall#
Wrapped from scipy.stats.crystalball; support: \(-\infty < z < \infty\); shape parameter(s): \(m > 1\), \(\beta > 0\).
Gibrat#
Wrapped from scipy.stats.gibrat; support: \(z > 0\).
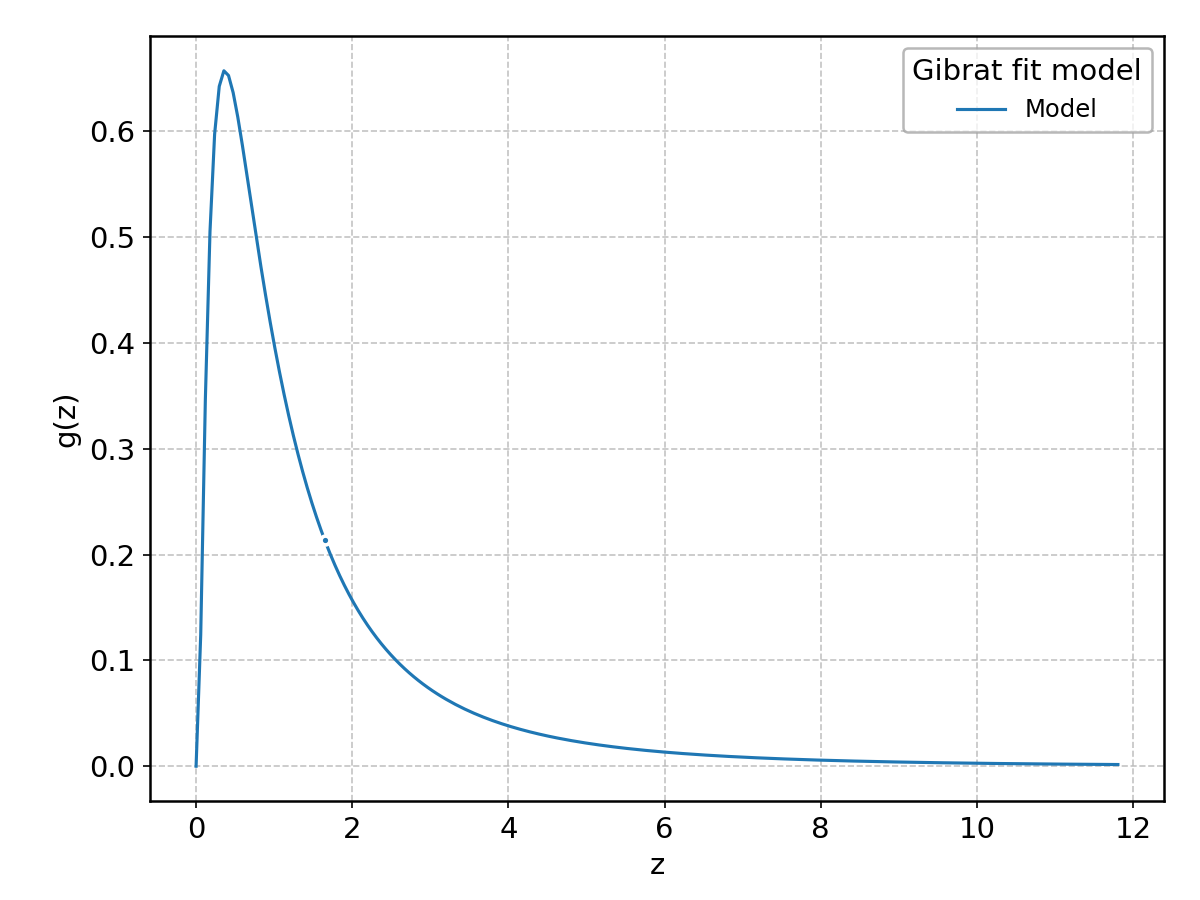
GumbelL#
Wrapped from scipy.stats.gumbel_l; support: \(-\infty < z < \infty\).
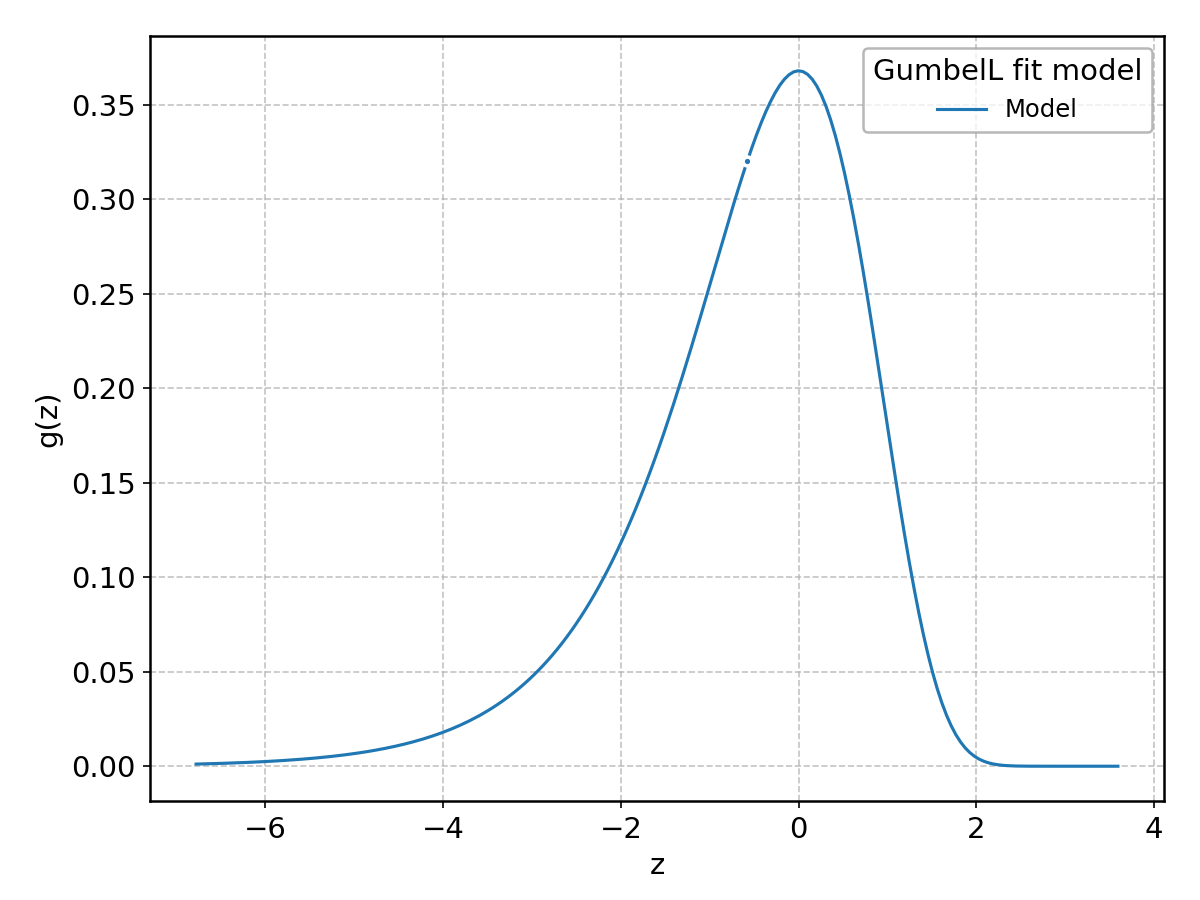
GumbelR#
Wrapped from scipy.stats.gumbel_r; support: \(-\infty < z < \infty\).
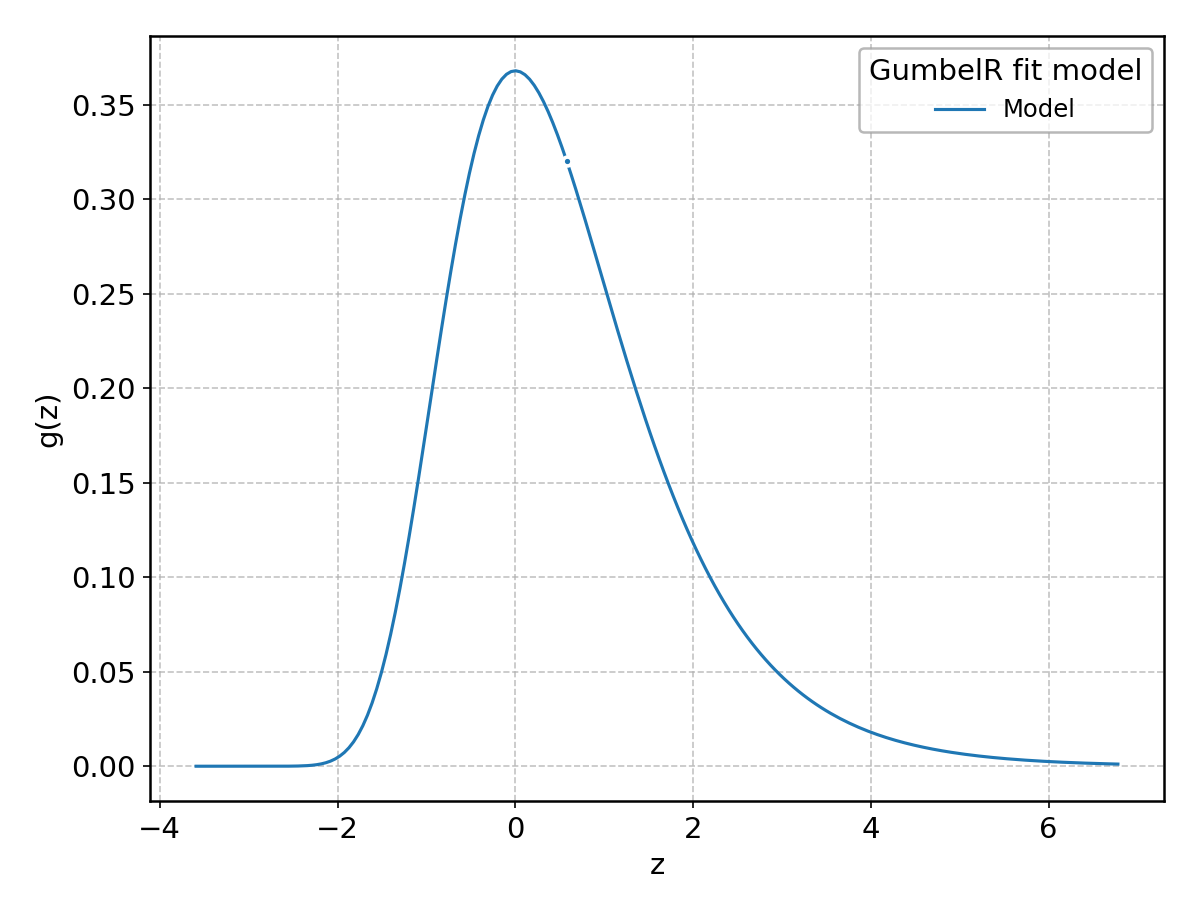
HalfCauchy#
Wrapped from scipy.stats.halfcauchy; support: \(z > 0\).
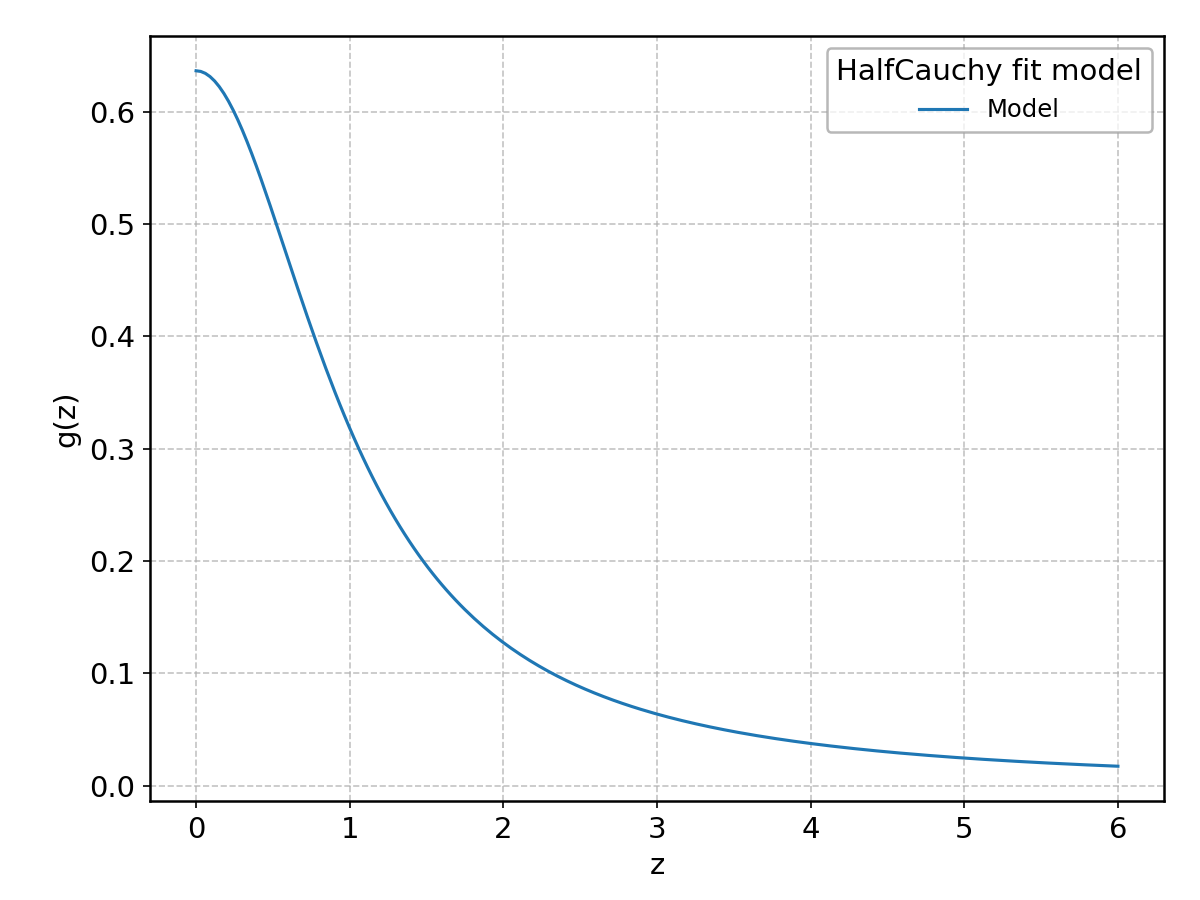
HalfLogistic#
Wrapped from scipy.stats.halflogistic; support: \(z > 0\).
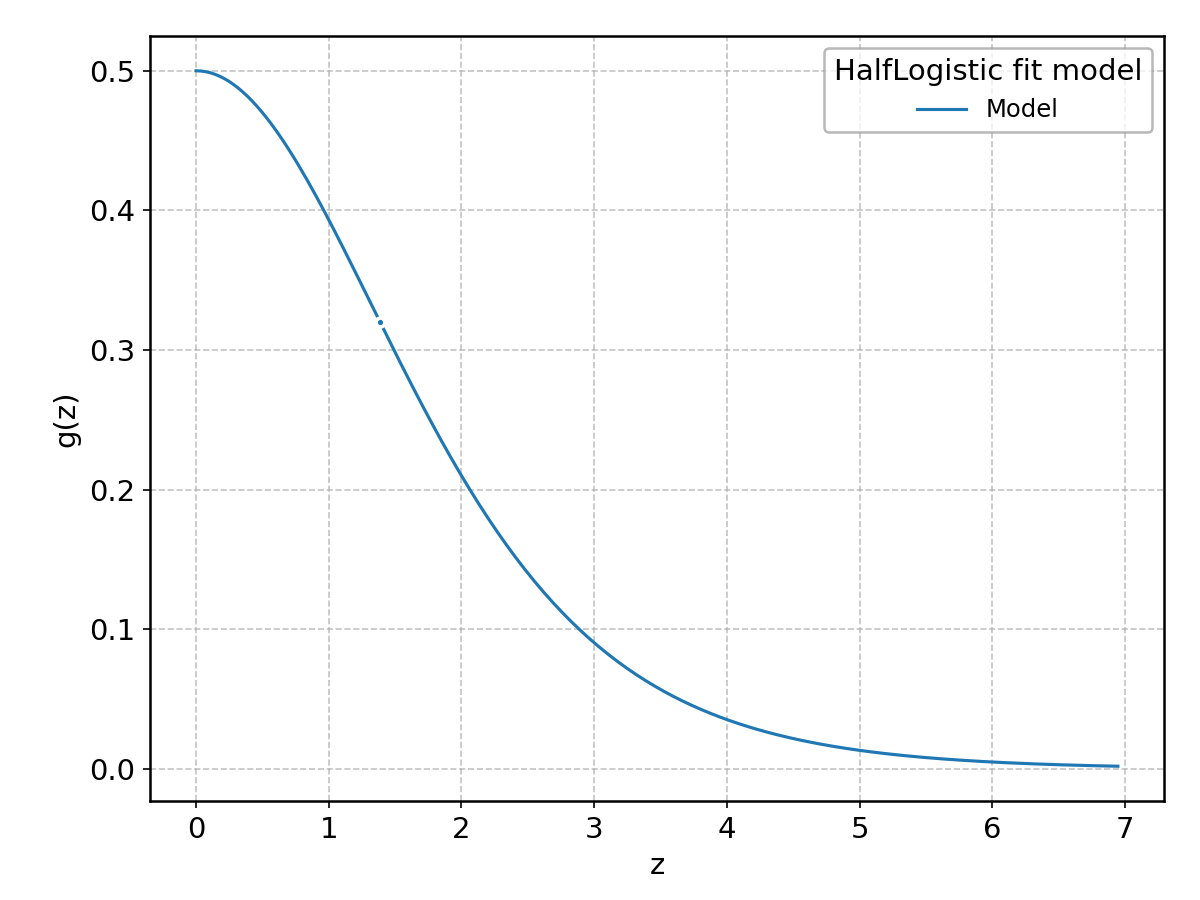
HalfNorm#
Wrapped from scipy.stats.halfnorm; support: \(z > 0\).
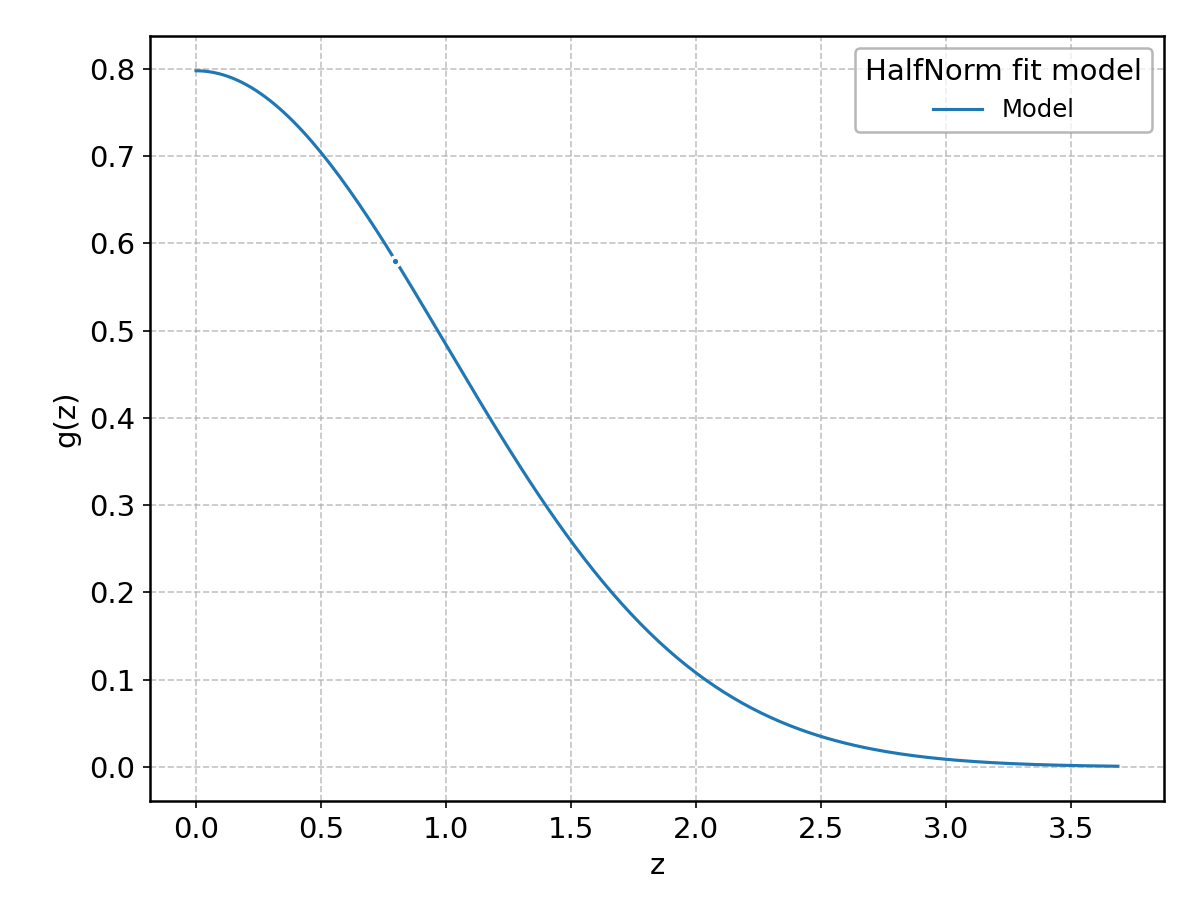
HyperSecant#
Wrapped from scipy.stats.hypsecant; support: \(-\infty < z < \infty\).
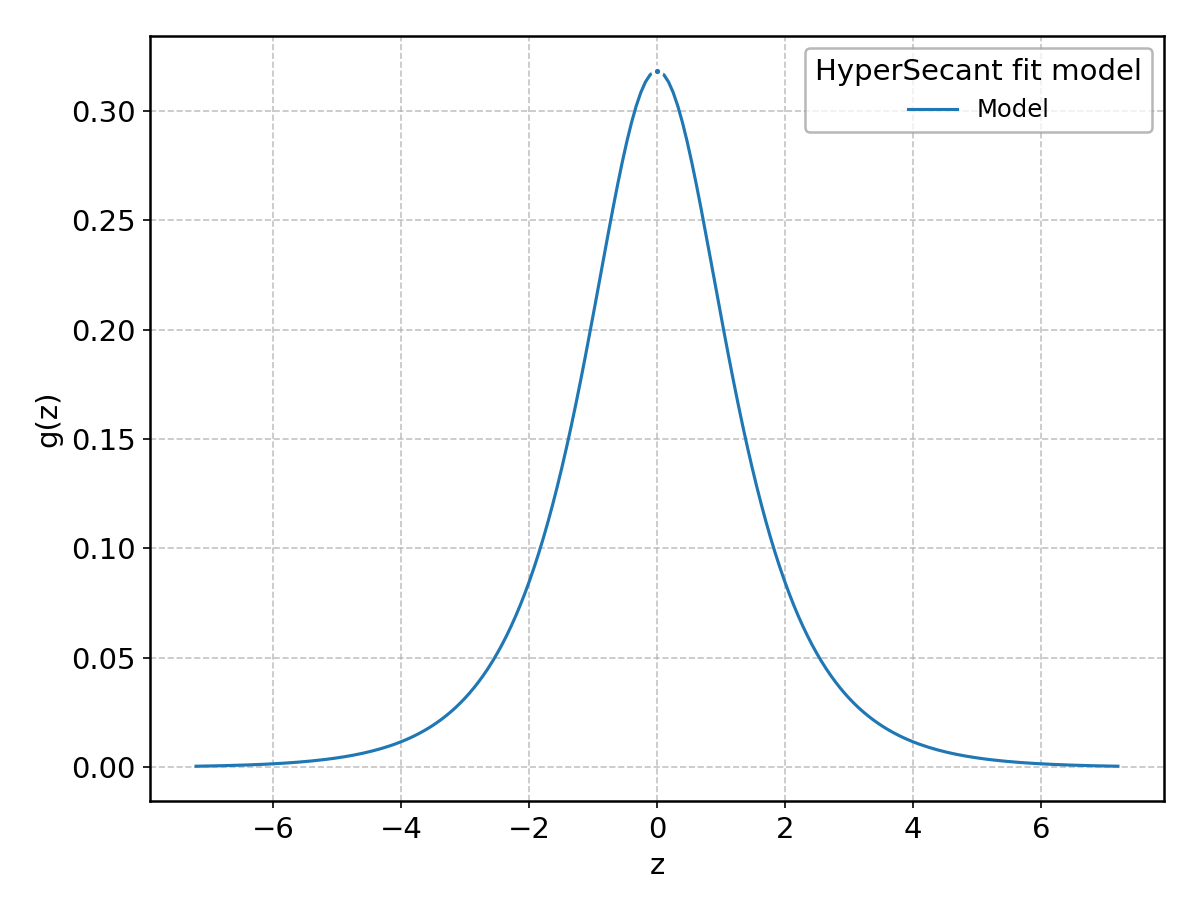
Landau#
Wrapped from scipy.stats.landau; support: \(-\infty < z < \infty\).
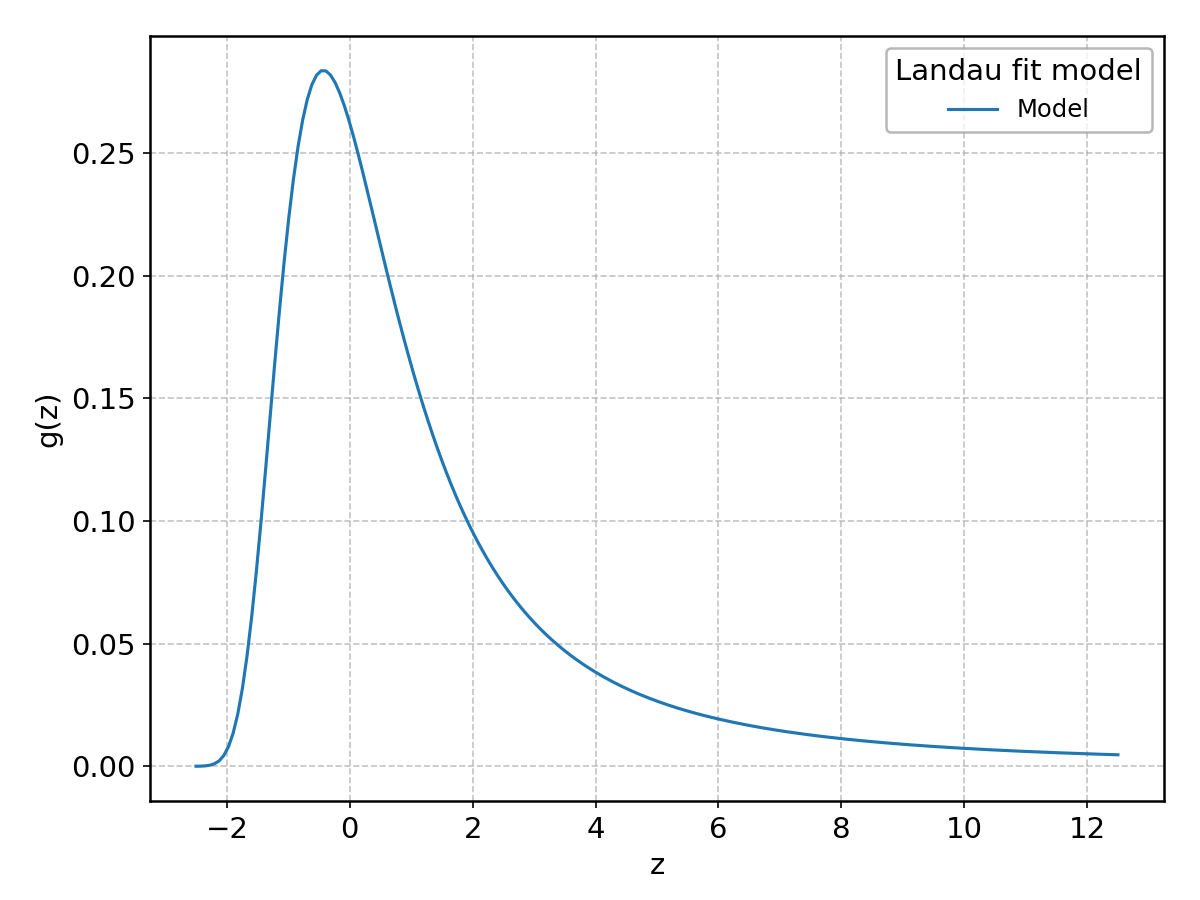
Laplace#
Wrapped from scipy.stats.laplace; support: \(-\infty < z < \infty\).
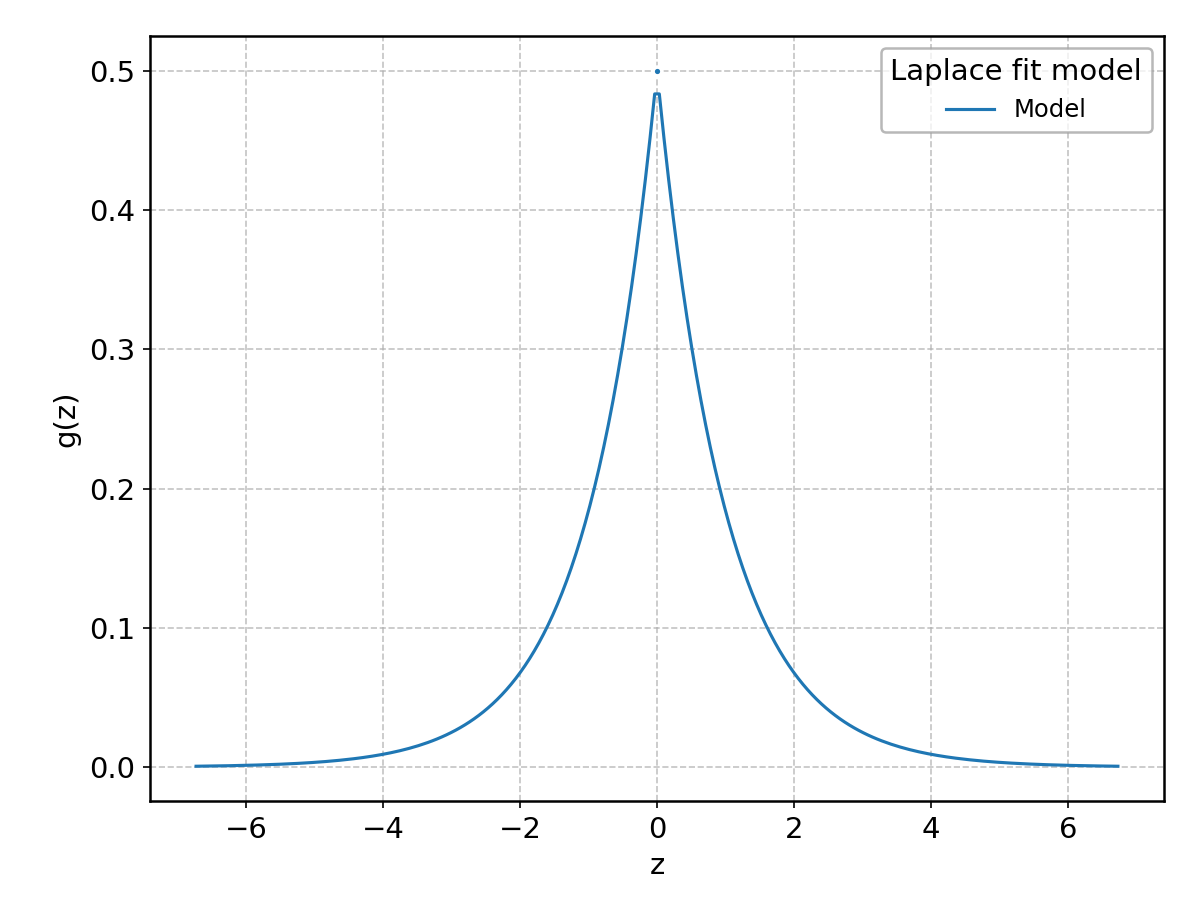
Levy#
Wrapped from scipy.stats.levy; support: \(-\infty < z < \infty\).
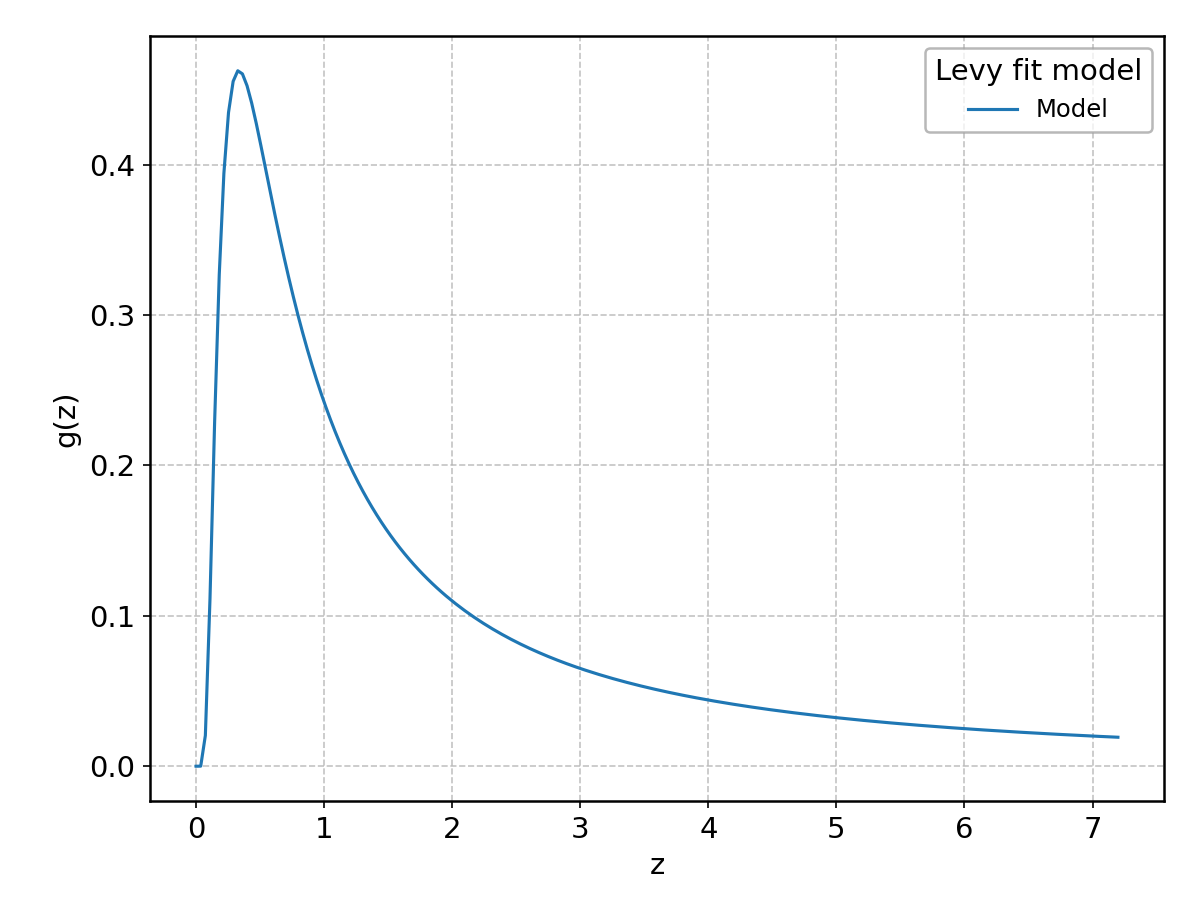
LevyL#
Wrapped from scipy.stats.levy_l; support: \(-\infty < z < \infty\).
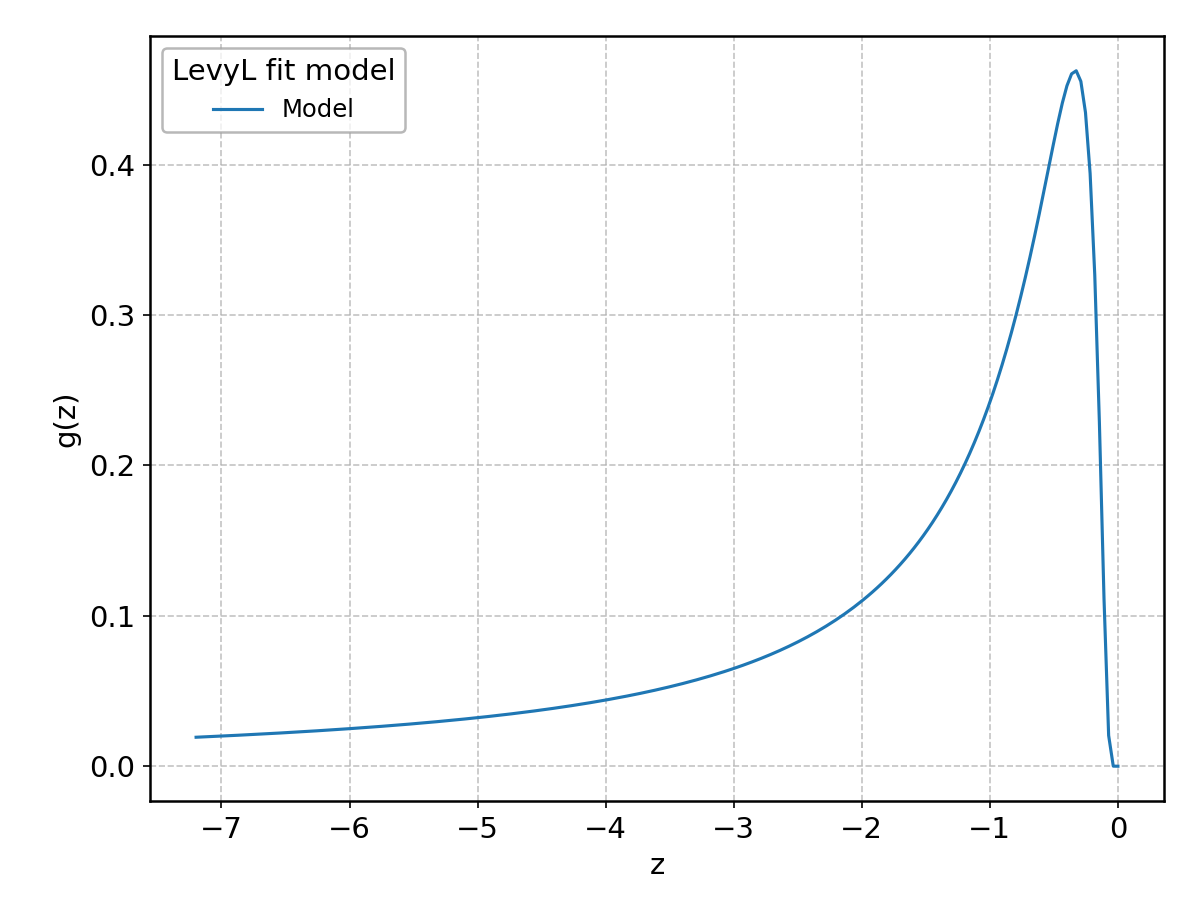
Logistic#
Wrapped from scipy.stats.logistic; support: \(-\infty < z < \infty\).
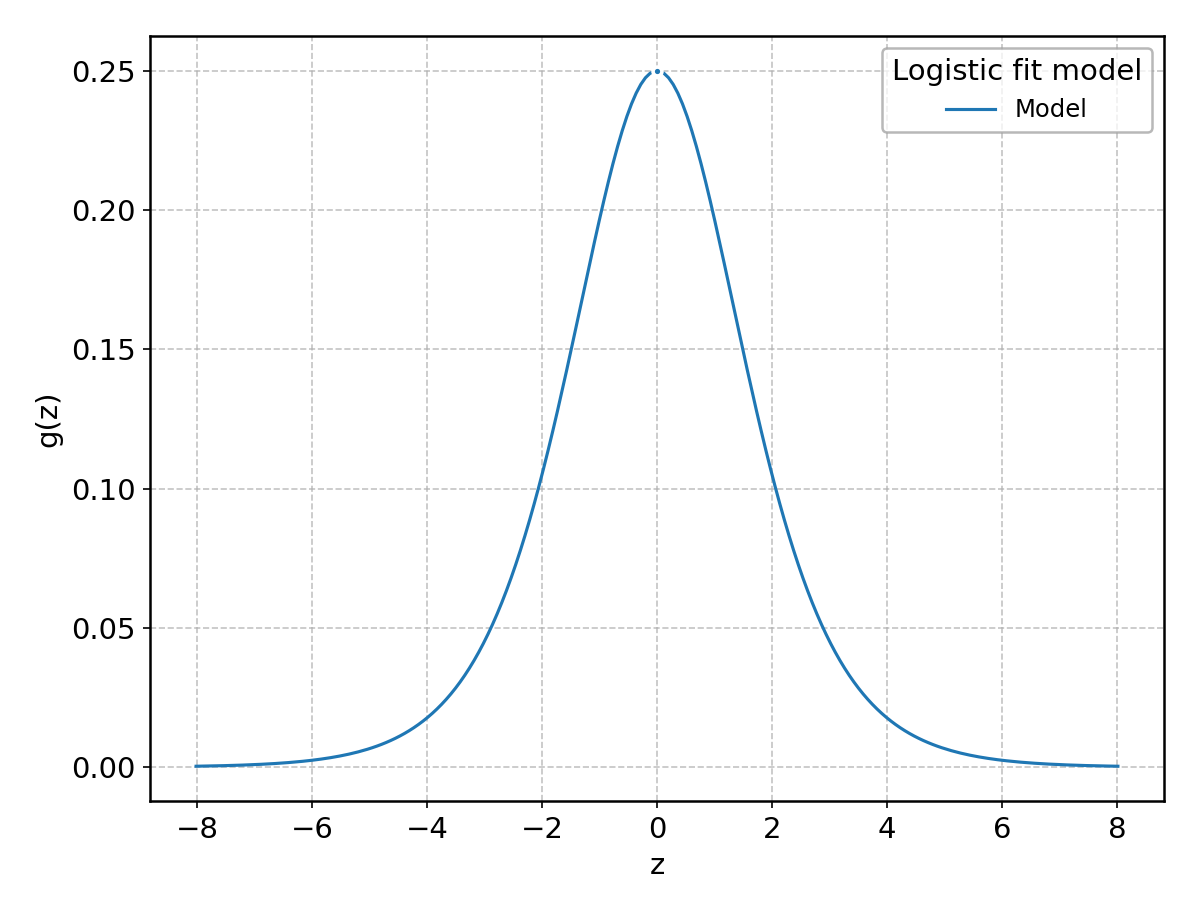
LogNormal#
Wrapped from scipy.stats.lognorm; support: \(z > 0\); shape parameter(s): \(s > 0\).
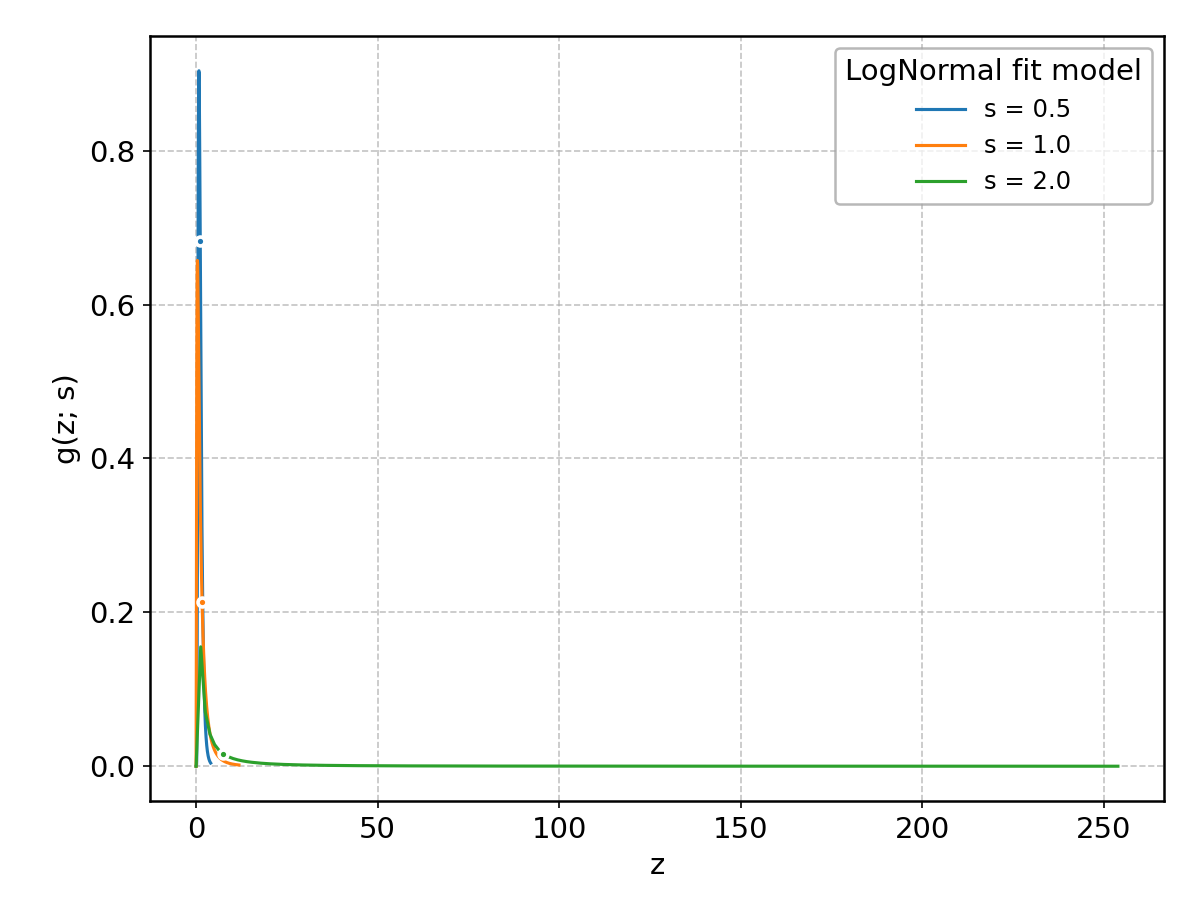
Maxwell#
Wrapped from scipy.stats.maxwell; support: \(0 < z < \infty\).
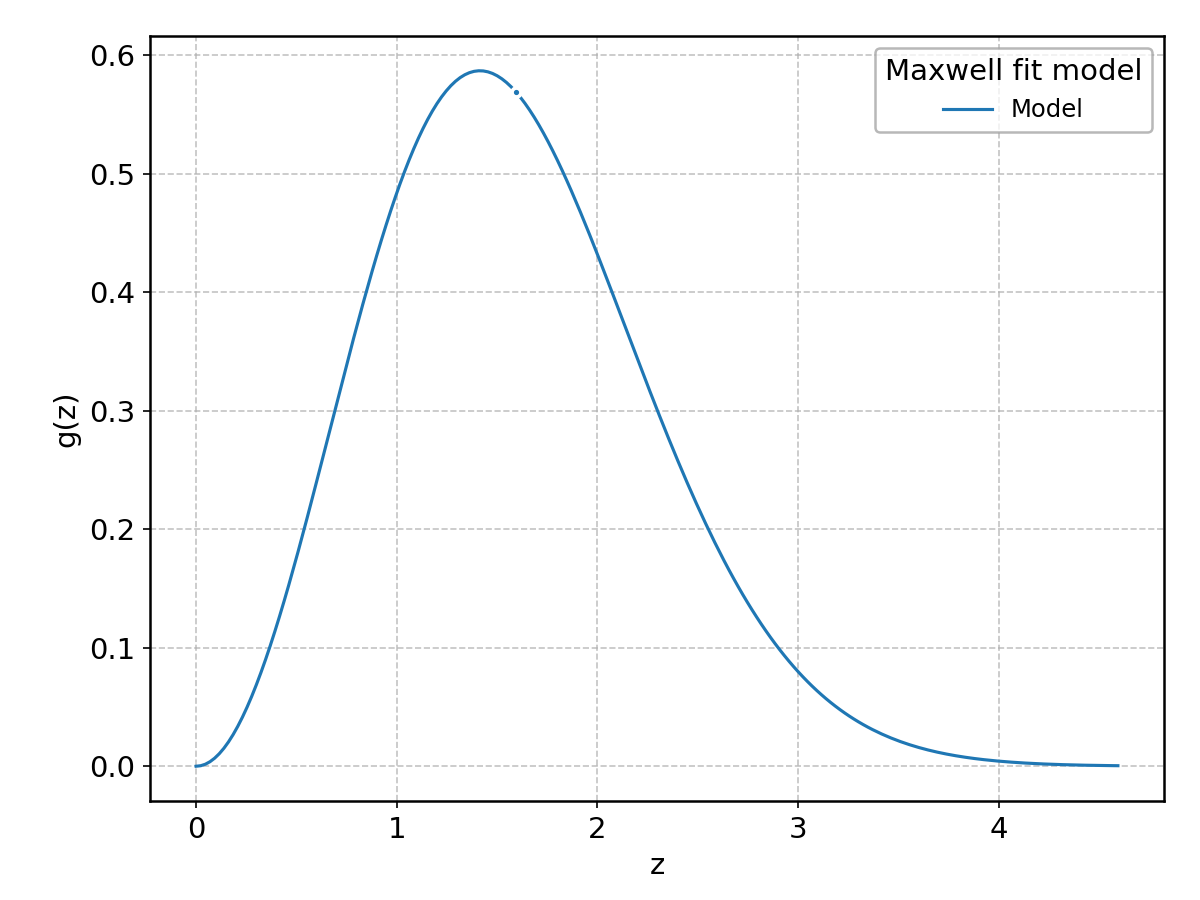
Moyal#
Wrapped from scipy.stats.moyal; support: \(-\infty < z < \infty\).
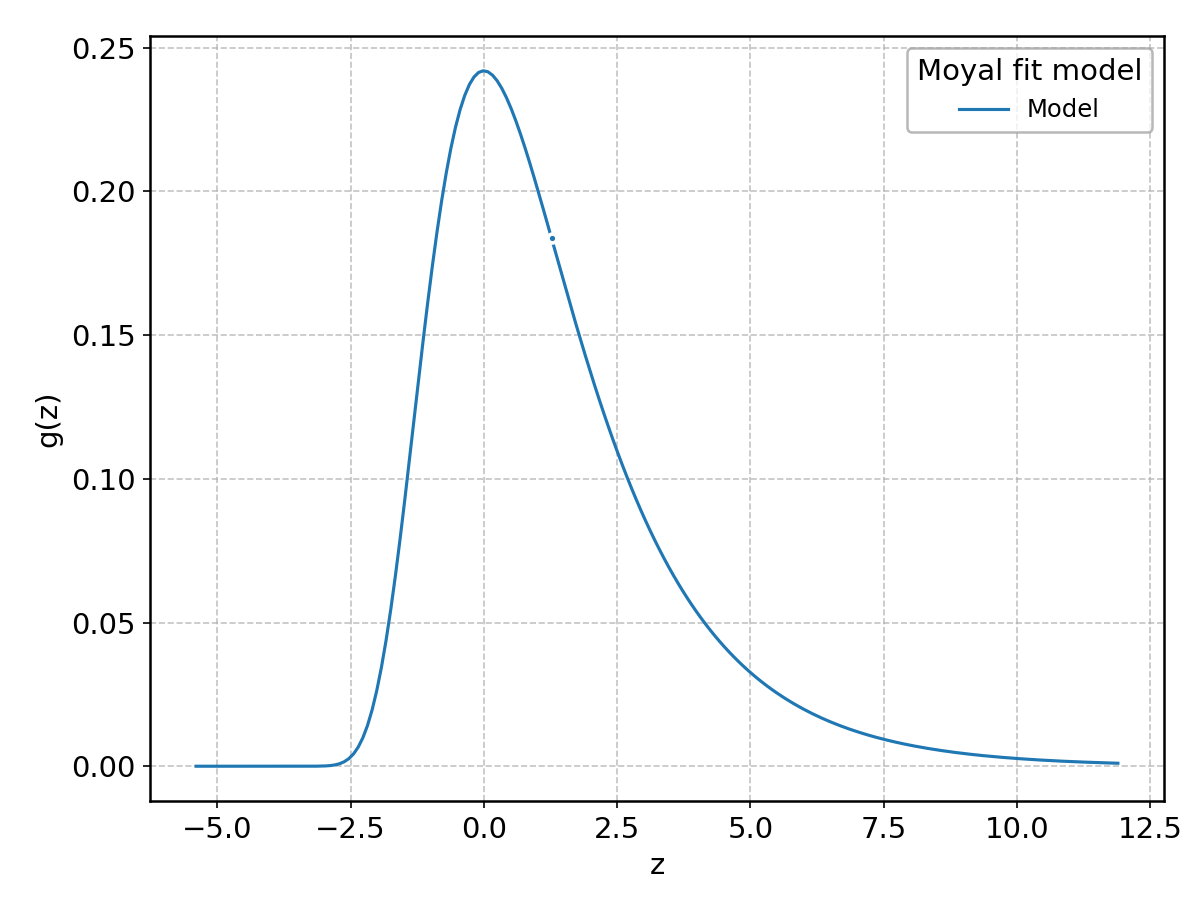
Nakagami#
Wrapped from scipy.stats.nakagami; support: \(0 < z < \infty\).
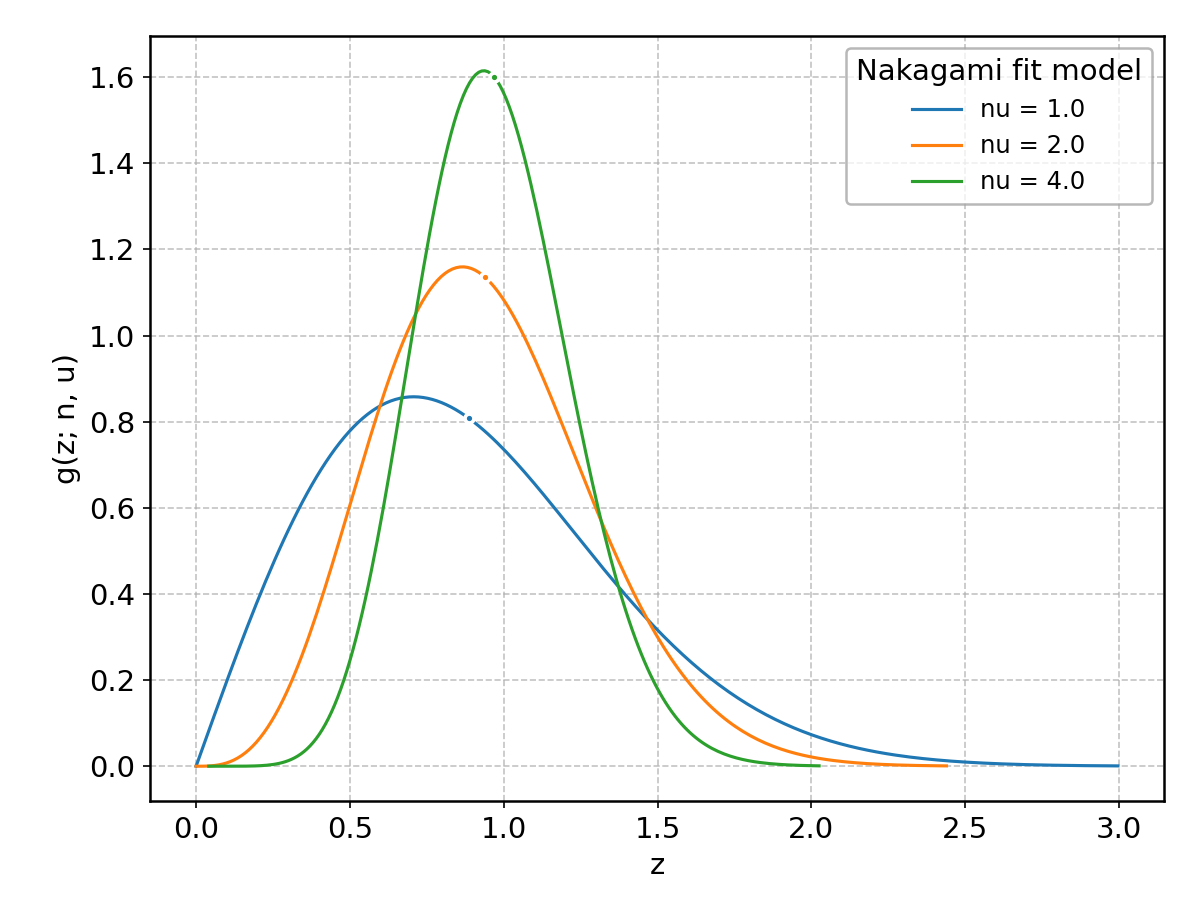
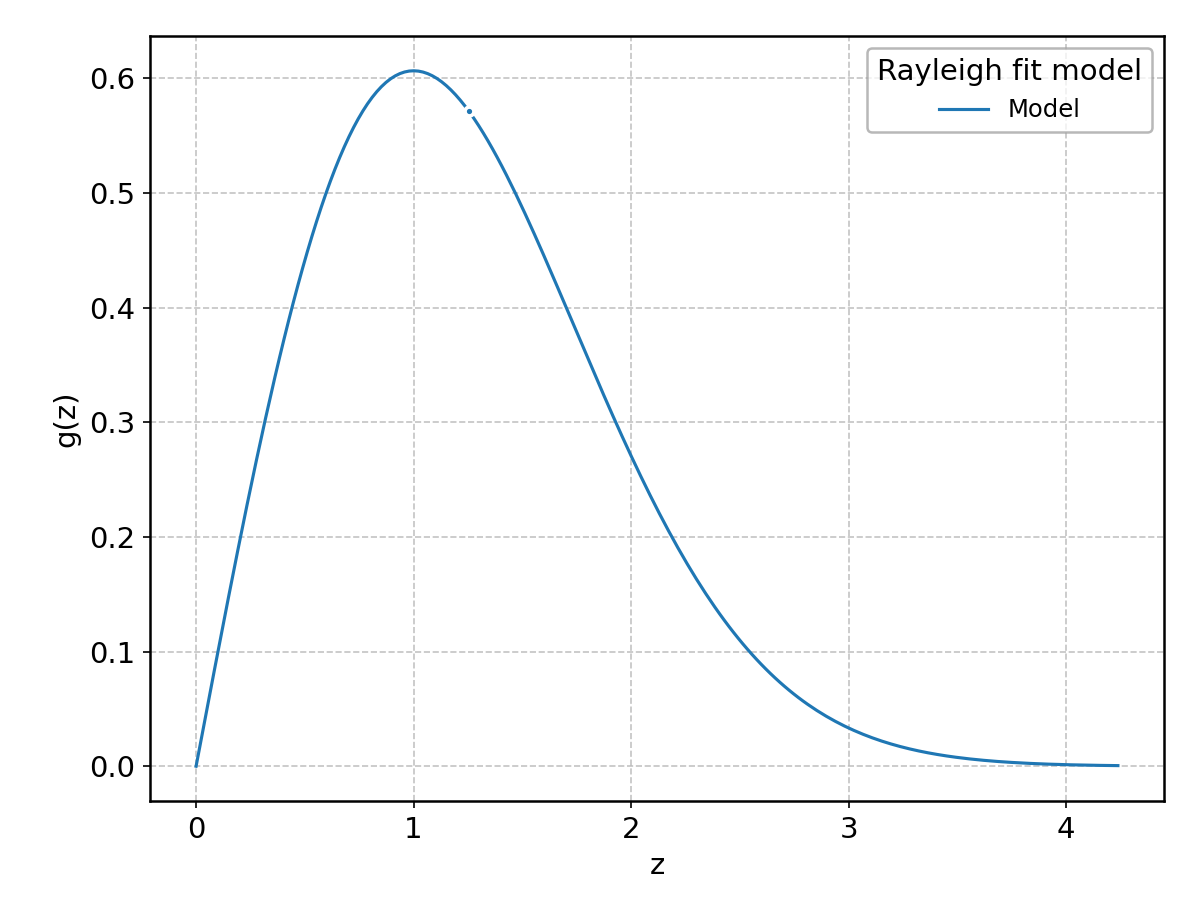
Rayleigh#
Wrapped from scipy.stats.rayleigh; support: \(0 < z < \infty\).
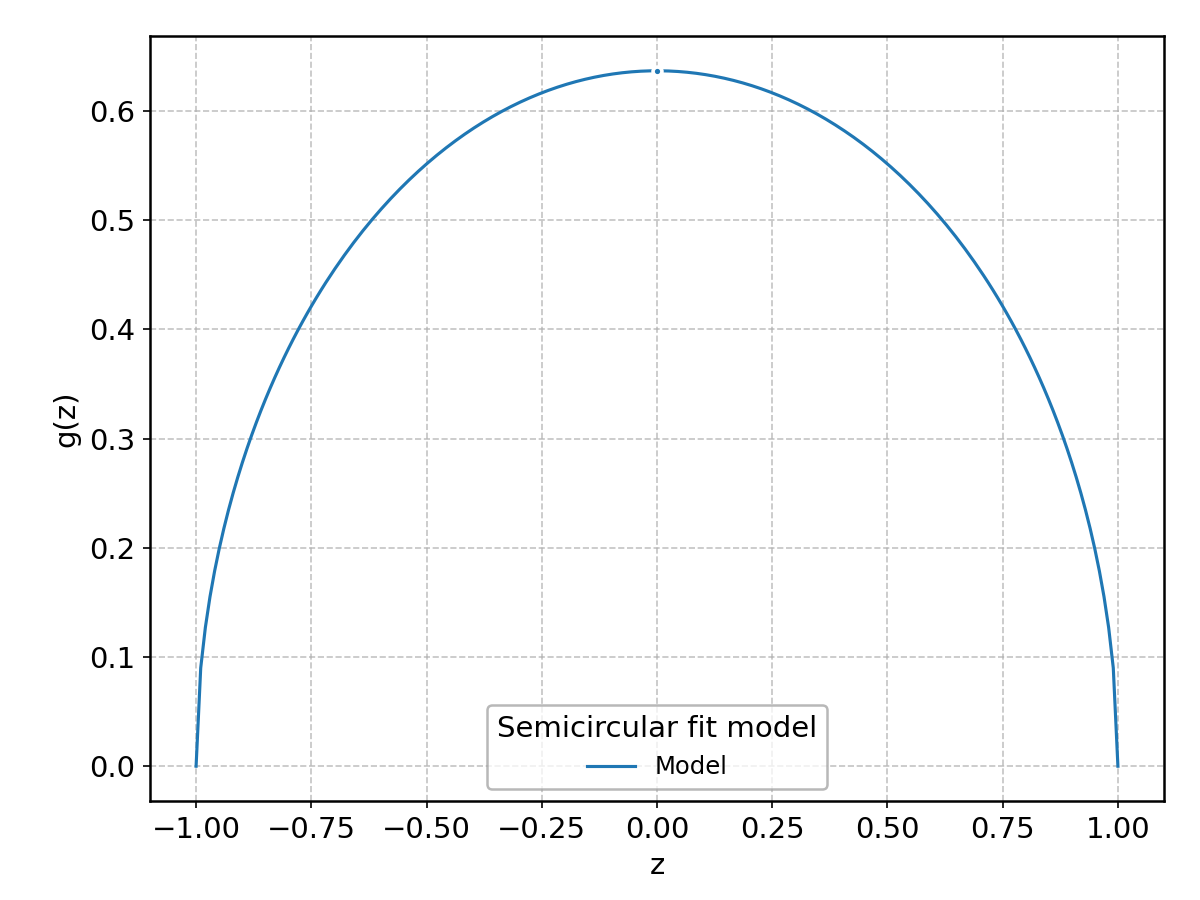
Semicircular#
Wrapped from scipy.stats.semicircular; support: \(-\infty < z < \infty\).
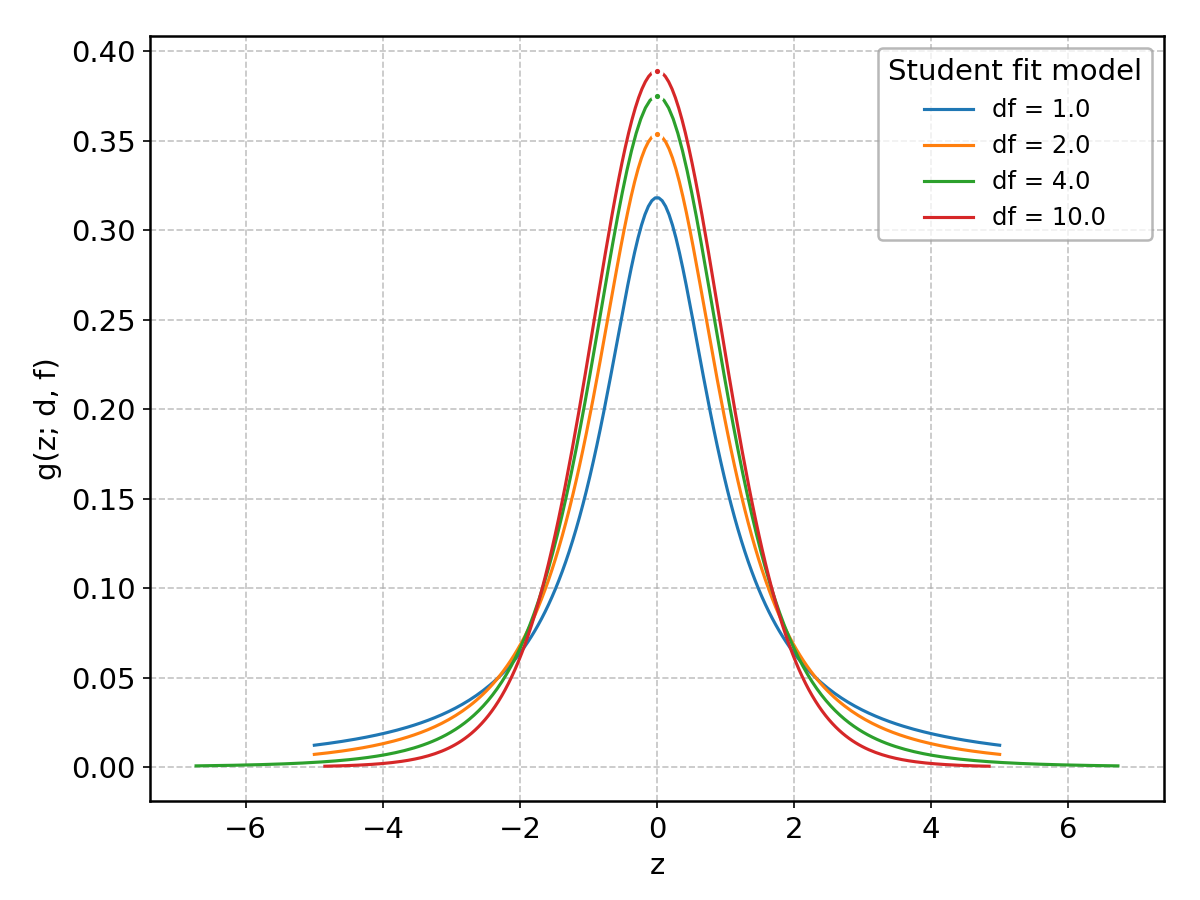
Student#
Wrapped from scipy.stats.t; support: \(-\infty < z < \infty\).
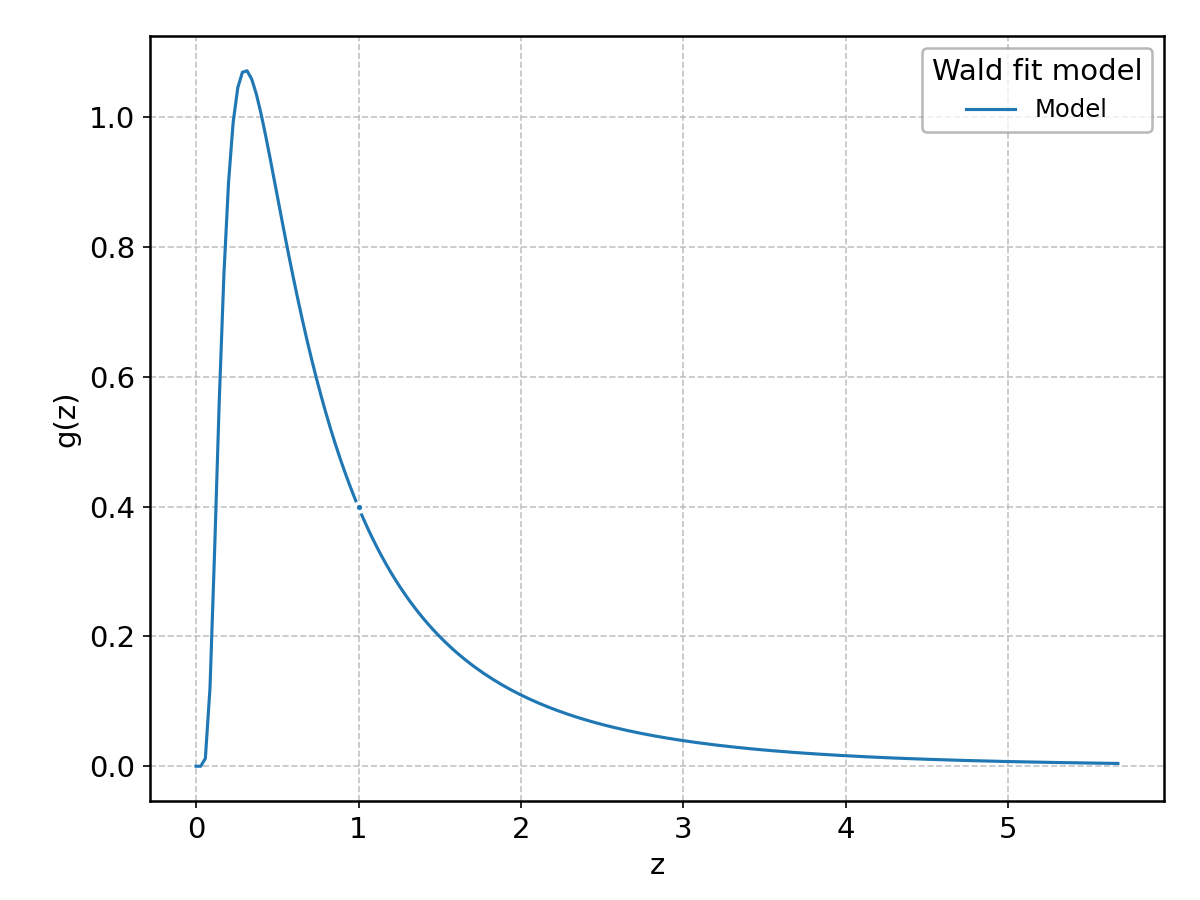
Wald#
Wrapped from scipy.stats.wald; support: \(0 < z < \infty\).
Module documentation#
Built in models.
- class aptapy.models.Constant(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Constant model.
- value = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- static evaluate(x: float | ndarray, value: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
This is simply using the weighted average of the y data, using the inverse of the squares of the errors as weights.
Note
This should provide the exact result in most cases, but, in the spirit of providing a common interface across all models, we are not overloading the fit() method. (Everything will continue working as expected, e.g., when one uses bounds on parameters.)
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Line(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Linear model.
- slope = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- intercept = FitParameter(value=0.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- static evaluate(x: float | ndarray, slope: float, intercept: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
This is simply using a weighted linear regression.
Note
This should provide the exact result in most cases, but, in the spirit of providing a common interface across all models, we are not overloading the fit() method. (Everything will continue working as expected, e.g., when one uses bounds on parameters.)
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Polynomial(degree: int, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Generic polynomial model.
Note that this is a convenience class to be used when one needs polynomials of arbitrary degree. For common low-order polynomials, consider using the dedicated classes (e.g., Line, Quadratic, etc.), which provide better initial parameter estimation.
Arguments#
- degreeint
The degree of the polynomial.
- labelstr, optional
The model label.
- xlabelstr, optional
The label for the x axis.
- ylabelstr, optional
The label for the y axis.
- static evaluate(x: float | ndarray, *coefficients: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
This is using a weighted linear regression.
Note
This should provide the exact result in most cases, but, in the spirit of providing a common interface across all models, we are not overloading the fit() method. (Everything will continue working as expected, e.g., when one uses bounds on parameters.)
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Quadratic(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Quadratic model.
This is just a convenience subclass of the generic Polynomial model with degree fixed to 2.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Cubic(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Cubic model.
This is just a convenience subclass of the generic Polynomial model with degree fixed to 3.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.PowerLaw(pivot: float = 1.0, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Power-law model.
Arguments#
- pivotfloat, optional
The pivot point of the power-law (default 1.).
- labelstr, optional
The model label.
- xlabelstr, optional
The label for the x axis.
- ylabelstr, optional
The label for the y axis.
- prefactor = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- index = FitParameter(value=-2.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- evaluate(x: float | ndarray, prefactor: float, index: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
This is using a weighted linear regression in log-log space. Note this is not an exact solution in the original space, for which a numerical optimization using non-linear least squares would be needed.
- default_plotting_range() Tuple[float, float][source]#
Overloaded method.
We might be smarter here, but for now we just return a fixed range that is not bogus when the index is negative, which should cover the most common use cases.
- plot(axes: Axes = None, fit_output: bool = False, **kwargs) None[source]#
Overloaded method.
In addition to the base class implementation, this also sets log scales on both axes.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Exponential(location: float = 0.0, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Exponential model.
Note this is an example of a model with a state, i.e., one where
evaluate()is not a static method, as we have alocationattribute that needs to be taken into account. This is done in the spirit of facilitating fits where the exponential decay starts at a non-zero x value.(One might argue that
locationshould be a fit parameter as well, but that would be degenerate with thescaleparameter, and it would have to be fixed in most cases anyway, so a simple attribute seems more appropriate here.)Arguments#
- locationfloat, optional
The location of the exponential decay (default 0.).
- labelstr, optional
The model label.
- xlabelstr, optional
The label for the x axis.
- ylabelstr, optional
The label for the y axis.
- prefactor = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- scale = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- evaluate(x: float | ndarray, prefactor: float, scale: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
This is using a weighted linear regression in lin-log space. Note this is not an exact solution in the original space, for which a numerical optimization using non-linear least squares would be needed.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.ExponentialComplement(location: float = 0.0, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Exponential complement model.
- evaluate(x: float | ndarray, prefactor: float, scale: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
Note we just pretend that the maximum of the y values is a reasonable estimate of the prefactor, and go back to the plain exponential case via the transformation ydata -> prefactor - ydata.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.StretchedExponential(location: float = 0.0, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Stretched exponential model.
- stretch = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- evaluate(x: float | ndarray, prefactor: float, scale: float, stretch: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0)[source]#
Overloaded method.
Note this a little bit flaky, in that we pretend that the data are well approximated by a plain exponential, and do not even try at estimating the stretch factor. When the latter is significantly different from 1 this will not be very accurate, but hopefully good enough to get the fit started.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.StretchedExponentialComplement(location: float = 0.0, label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Stretched exponential complement model.
- evaluate(x: float | ndarray, prefactor: float, scale: float, stretch: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
See the comment in the corresponding docstrings of the ExponentialComplement class.
- _abc_impl = <_abc._abc_data object>#
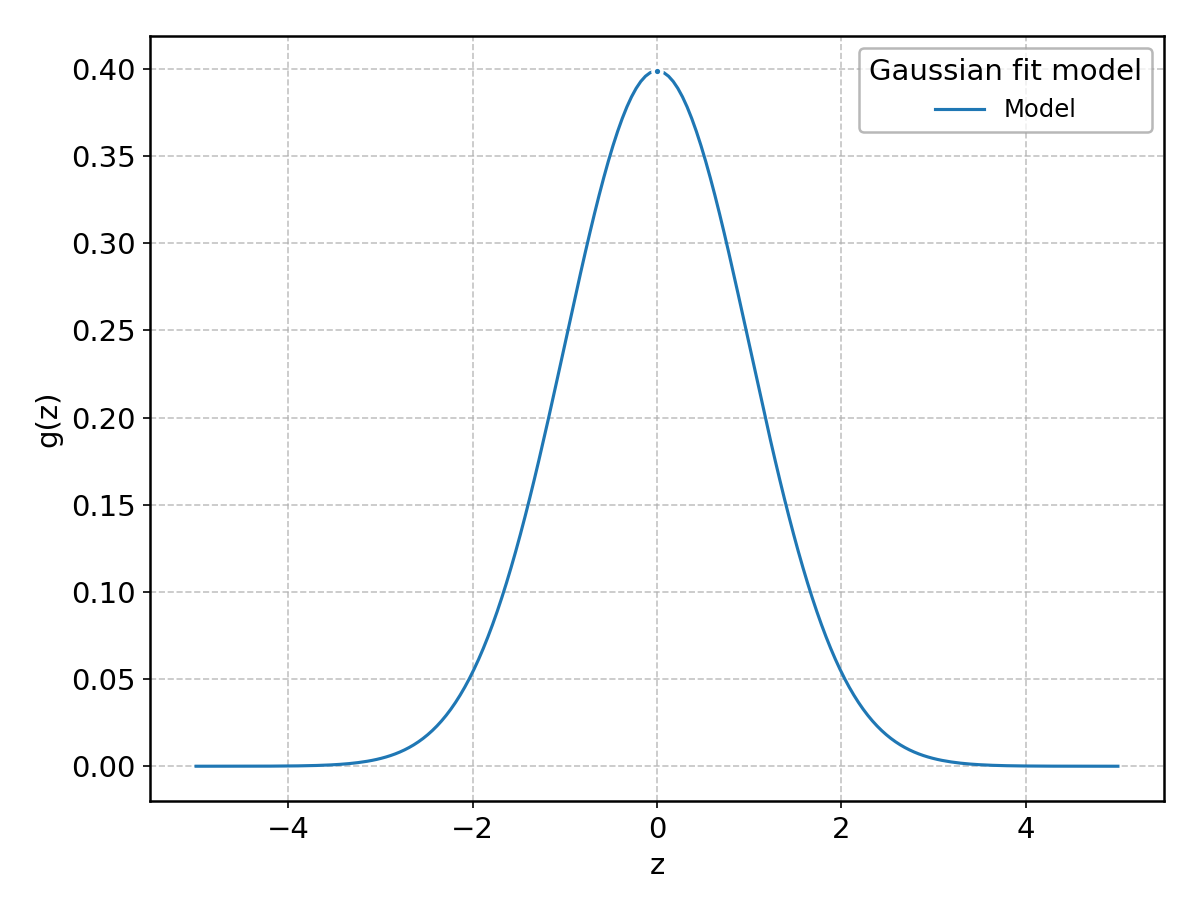
- class aptapy.models.Gaussian(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
This is a re-implementation from scratch of the normal distribution, which is also available as a wrapper around scipy.stats.norm as Normal.
The main reason for this is to be able to call the underlying parameters with more familiar names (mu, sigma, rather than loc, scale), as well as providing additional convenience methods such the iterative fitting around the peak.
- amplitude = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- mu = FitParameter(value=0.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- sigma = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- static evaluate(x, amplitude, mu, sigma, *args)[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- rvs(size: int = 1, random_state=None)[source]#
Generate random variates from the underlying distribution at the current parameter values.
Arguments#
- sizeint, optional
The number of random variates to generate (default 1).
- random_stateint or np.random.Generator, optional
The random seed or generator to use (default None).
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
- fit_iterative(xdata: float | ndarray | Histogram1d, ydata: float | ndarray = None, *, p0: float | ndarray = None, sigma: float | ndarray = None, num_sigma_left: float = 2.0, num_sigma_right: float = 2.0, num_iterations: int = 2, **kwargs) FitStatus[source]#
Fit the core of Gaussian data within a given number of sigma around the peak.
This function performs a first round of fit to the data (either a histogram or scatter plot data) and then repeats the fit iteratively, limiting the fit range to a specified interval defined in terms of deviations (in sigma) around the peak.
Arguments#
- xdataarray_like or Histogram1d
The data (scatter plot x values) or histogram to fit.
- ydataarray_like, optional
The y data to fit (if xdata is not a Histogram1d).
- p0array_like, optional
The initial values for the fit parameters.
- sigmaarray_like, optional
The uncertainties on the y data.
- num_sigma_leftfloat
The number of sigma on the left of the peak to be used to define the fitting range.
- num_sigma_rightfloat
The number of sigma on the right of the peak to be used to define the fitting range.
- num_iterationsint
The number of iterations of the fit.
- kwargsdict, optional
Additional keyword arguments passed to fit().
Returns#
- FitStatus
The results of the fit.
- plot(axes: Axes = None, fit_output: bool = False, plot_mean: bool = True, **kwargs) None[source]#
Plot the model.
Note this is reimplemented from scratch to allow overplotting the mean of the distribution.
Arguments#
- axesmatplotlib.axes.Axes, optional
The axes to plot on (default: current axes).
- fit_outputbool, optional
Whether to include the fit output in the legend (default: False).
- plot_meanbool, optional
Whether to overplot the mean of the distribution (default: True).
- kwargsdict, optional
Additional keyword arguments passed to plt.plot().
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Fe55Forest(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Model representing the Kα and Kβ emission lines produced in the decay of 55Fe. The energy values are computed as the intensity-weighted mean of all possible emission lines contributing to each feature.
The energy data are retrieved from the X-ray database at https://xraydb.seescience.org/
- TABULATED_KB_INTENSITY = 0.12445#
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
- _abc_impl = <_abc._abc_data object>#
- amplitude = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- energies = (5.896, 6.492)#
- energy_scale = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- intensity1 = FitParameter(value=0.5, _name=None, error=None, _frozen=False, minimum=0.0, maximum=1.0)#
- sigma = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Probit(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Custom implementation of the probit model, i.e., the percent-point function of a gaussian distribution.
- offset = FitParameter(value=0.0, _name=None, error=None, _frozen=False, minimum=-inf, maximum=inf)#
- sigma = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- static evaluate(x: float | ndarray, offset: float, sigma: float) float | ndarray[source]#
Evaluate the model at a given set of parameter values.
Arguments#
- xarray_like
The value(s) of the independent variable.
- parameter_valuessequence of float
The value of the model parameters.
Returns#
- yarray_like
The value(s) of the model at the given value(s) of the independent variable for a given set of parameter values.
- init_parameters(xdata: float | ndarray, ydata: float | ndarray, sigma: float | ndarray = 1.0) None[source]#
Overloaded method.
- static default_plotting_range() Tuple[float, float][source]#
Overloaded method.
Since the probit function diverges at 0 and 1, we limit the plotting range to a reasonable interval.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.ErfSigmoid(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Error function model.
- static shape(z)[source]#
Abstract method for the normalized shape of the sigmoid model. Subclasses must implement this method.
Arguments#
- zarray_like
The normalized independent variable.
- parameter_valuesfloat
Additional shape parameters for the sigmoid.
Returns#
- array_like
The value of the sigmoid shape function at z.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.LogisticSigmoid(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Logistic function model.
- static shape(z)[source]#
Abstract method for the normalized shape of the sigmoid model. Subclasses must implement this method.
Arguments#
- zarray_like
The normalized independent variable.
- parameter_valuesfloat
Additional shape parameters for the sigmoid.
Returns#
- array_like
The value of the sigmoid shape function at z.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Arctangent(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Arctangent function model.
- static shape(z)[source]#
Abstract method for the normalized shape of the sigmoid model. Subclasses must implement this method.
Arguments#
- zarray_like
The normalized independent variable.
- parameter_valuesfloat
Additional shape parameters for the sigmoid.
Returns#
- array_like
The value of the sigmoid shape function at z.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.HyperbolicTangent(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Hyperbolic tangent function model.
- static shape(z)[source]#
Abstract method for the normalized shape of the sigmoid model. Subclasses must implement this method.
Arguments#
- zarray_like
The normalized independent variable.
- parameter_valuesfloat
Additional shape parameters for the sigmoid.
Returns#
- array_like
The value of the sigmoid shape function at z.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Alpha(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.alpha_gen object>#
- a = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Anglit(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.anglit_gen object>#
- class aptapy.models.Arcsine(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.arcsine_gen object>#
- class aptapy.models.Argus(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.argus_gen object>#
- chi = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Beta(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.beta_gen object>#
- a = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- b = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.BetaPrime(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.betaprime_gen object>#
- a = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- b = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Bradford(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.bradford_gen object>#
- c = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Burr(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.burr_gen object>#
- c = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- d = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Burr12(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.burr12_gen object>#
- c = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- d = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Cauchy(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.cauchy_gen object>#
- class aptapy.models.Chi(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.chi_gen object>#
- df = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Chisquare(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.chi2_gen object>#
- df = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Cosine(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.cosine_gen object>#
- class aptapy.models.CrystalBall(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.crystalball_gen object>#
- beta = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- m = FitParameter(value=2.0, _name=None, error=None, _frozen=False, minimum=1.0, maximum=inf)#
- class aptapy.models.Gibrat(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.gibrat_gen object>#
- class aptapy.models.GumbelL(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.gumbel_l_gen object>#
- class aptapy.models.GumbelR(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.gumbel_r_gen object>#
- class aptapy.models.HalfCauchy(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.halfcauchy_gen object>#
- class aptapy.models.HalfLogistic(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.halflogistic_gen object>#
- class aptapy.models.HalfNorm(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.halfnorm_gen object>#
- class aptapy.models.HyperSecant(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.hypsecant_gen object>#
- class aptapy.models.Landau(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- default_plotting_range() Tuple[float, float][source]#
Overloaded method.
The Landau distribution is peculiar in that it has no definite mean or variance, and its support is unbounded. It is also asymmetric, with a long right tail. Therefore, we resort to a custom function for the plotting range.
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.landau_gen object>#
- class aptapy.models.Laplace(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.laplace_gen object>#
- class aptapy.models.Levy(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.levy_gen object>#
- class aptapy.models.LevyL(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.levy_l_gen object>#
- class aptapy.models.Logistic(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.logistic_gen object>#
- class aptapy.models.LogNormal(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.lognorm_gen object>#
- s = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Lorentzian(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
Alias for the Cauchy distribution.
- _abc_impl = <_abc._abc_data object>#
- class aptapy.models.Maxwell(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.maxwell_gen object>#
- class aptapy.models.Moyal(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.moyal_gen object>#
- class aptapy.models.Nakagami(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.nakagami_gen object>#
- nu = FitParameter(value=1.0, _name=None, error=None, _frozen=False, minimum=0.0, maximum=inf)#
- class aptapy.models.Normal(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.norm_gen object>#
- class aptapy.models.Rayleigh(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.rayleigh_gen object>#
- class aptapy.models.Semicircular(label: str = None, xlabel: str = None, ylabel: str = None)[source]#
- _abc_impl = <_abc._abc_data object>#
- _rv = <scipy.stats._continuous_distns.semicircular_gen object>#